Review Article |
|
Corresponding author: Jacquelyn C. Guzy ( jguzy@usgs.gov ) Academic editor: Sven Bacher
© 2023 Jacquelyn C. Guzy, Bryan G. Falk, Brian J. Smith, John David Willson, Robert N. Reed, Nicholas G. Aumen, Michael L. Avery, Ian A. Bartoszek, Earl Campbell, Michael S. Cherkiss, Natalie M. Claunch, Andrea F. Currylow, Tylan Dean, Jeremy Dixon, Richard Engeman, Sarah Funck, Rebekah Gibble, Kodiak C. Hengstebeck, John S. Humphrey, Margaret E. Hunter, Jillian M. Josimovich, Jennifer Ketterlin, Michael Kirkland, Frank J. Mazzotti, Robert McCleery, Melissa A. Miller, Matthew McCollister, M. Rockwell Parker, Shannon E. Pittman, Michael Rochford, Christina Romagosa, Art Roybal, Ray W. Snow, McKayla M. Spencer, J. Hardin Waddle, Amy A. Yackel Adams, Kristen M. Hart.
This is an open access article distributed under the terms of the CC0 Public Domain Dedication.
Citation:
Guzy JC, Falk BG, Smith BJ, Willson JD, Reed RN, Aumen NG, Avery ML, Bartoszek IA, Campbell E, Cherkiss MS, Claunch NM, Currylow AF, Dean T, Dixon J, Engeman R, Funck S, Gibble R, Hengstebeck KC, Humphrey JS, Hunter ME, Josimovich JM, Ketterlin J, Kirkland M, Mazzotti FJ, McCleery R, Miller MA, McCollister M, Parker MR, Pittman SE, Rochford M, Romagosa C, Roybal A, Snow RW, Spencer MM, Waddle JH, Yackel Adams AA, Hart KM (2023) Burmese pythons in Florida: A synthesis of biology, impacts, and management tools. NeoBiota 80: 1-119. https://doi.org/10.3897/neobiota.80.90439
|
Abstract
Burmese pythons (Python molurus bivittatus) are native to southeastern Asia, however, there is an established invasive population inhabiting much of southern Florida throughout the Greater Everglades Ecosystem. Pythons have severely impacted native species and ecosystems in Florida and represent one of the most intractable invasive-species management issues across the globe. The difficulty stems from a unique combination of inaccessible habitat and the cryptic and resilient nature of pythons that thrive in the subtropical environment of southern Florida, rendering them extremely challenging to detect. Here we provide a comprehensive review and synthesis of the science relevant to managing invasive Burmese pythons. We describe existing control tools and review challenges to productive research, identifying key knowledge gaps that would improve future research and decision making for python control.
Keywords
control tools, demography, detection, Florida, impacts, invasive species, Python molurus bivittatus, reptile, suppression
Introduction
Burmese pythons (Python molurus bivittatus, see Taxonomy section) are oviparous (egg-laying), nonvenomous, large, constricting snakes native to southeastern Asia (
The Greater Everglades Ecosystem consists of a vast, shallow, watershed 87 km (60 miles) long and 161 km (100 miles) wide (
General Burmese python (Python molurus bivittatus) research areas across southern Florida (inset, black box). Primary research areas are indicated by green shaded polygons or black dots. Large black dots indicate major cities and gray lines indicate major roads; the beige line indicates Loop Road (unpaved). Levee-67 continues into ENP (L-67X). Abbreviations include Arthur R. Marshall Loxahatchee National Wildlife Refuge (LNWR), Everglades and Francis S. Taylor Wildlife Management Area (Everglades WMA), Water Conservation Areas (WCA 2 and 3), Stormwater Treatment Area 3/4 (STA 3/4), Pa-hay-okee Road (Pa-hay-okee), Hole-in-the-Donut Restoration Area (Hole-in-the-Donut), Frog Pond Wildlife Management Area (Frog Pond WMA), Rookery Bay National Estuarine Research Reserve (RBNERR), and Crocodile Lake National Wildlife Refuge (CLNWR). Faint gray lines are county boundaries.
Locations or entities referenced in this manuscript and the corresponding acronym.
| Location, Agency, or Term | Acronym |
|---|---|
| Arthur R. Marshall Loxahatchee National Wildlife Refuge | LNWR |
| Big Cypress National Preserve | BICY |
| Biscayne National Park | BISC |
| Close-kin mark-recapture | CKMR |
| Collier-Seminole State Park | CSSP |
| Convention on International Trade in Endangered Species of Wild Fauna and Flora | CITES |
| Conservancy of Southwest Florida | CSWFL |
| Crocodile Lake National Wildlife Refuge | CLNWR |
| Department of Interior | DOI |
| Everglades and Francis S. Taylor Wildlife Management Area | Everglades WMA |
| Everglades National Park | ENP |
| Fakahatchee Strand Preserve State Park | FSPSP |
| Florida Department of Environmental Protection | DEP |
| Florida Fish and Wildlife Conservation Commission | FWC |
| Frog Pond Wildlife Management Area | Frog Pond WMA |
| Hole-in-the-Donut Restoration Area | Hole-in-the-Donut |
| International Commission on Zoological Nomenclature | ICZN |
| National Park Service | NPS |
| Pa-hay-okee Road | Pa-hay-okee |
| Picayune Strand State Forest | PSSF |
| Rookery Bay National Estuarine Research Reserve | RBNERR |
| South Florida Water Management District | SFWMD |
| Stormwater Treatment Area | STA |
| United States Fish and Wildlife Service | USFWS |
| United States Geological Survey | USGS |
| United States Department of Agriculture | USDA |
| Water Conservation Area | WCA |
Although Burmese pythons were found in the Greater Everglades as early as 1979 (
In this synthesis we discuss Burmese python biology as it relates to management of the species, including insights from research on their taxonomy, demography, and physiology. Second, we outline our current understanding of how Burmese pythons arrived in Florida. Third, we review one of the greatest challenges in managing Burmese pythons, low detectability, and progress addressing this challenge. We then discuss ecological and socioeconomic impacts of pythons and describe the distribution and movement ecology of the species. Finally, we describe existing control tools and review challenges to productive research, identifying key knowledge gaps that could improve future research and decision making for python control.
The authorship of this publication is diverse, representing many of the experienced scientists and managers involved with the Burmese python invasion over the last two decades, including representatives from United States federal agencies, the state of Florida, and numerous non-profit and academic institutions. This document represents the consensus of this scientific community, and in the few cases where consensus is not clear, multiple viewpoints are explored for better insight to drive future research.
Natural history of Burmese pythons
Identification
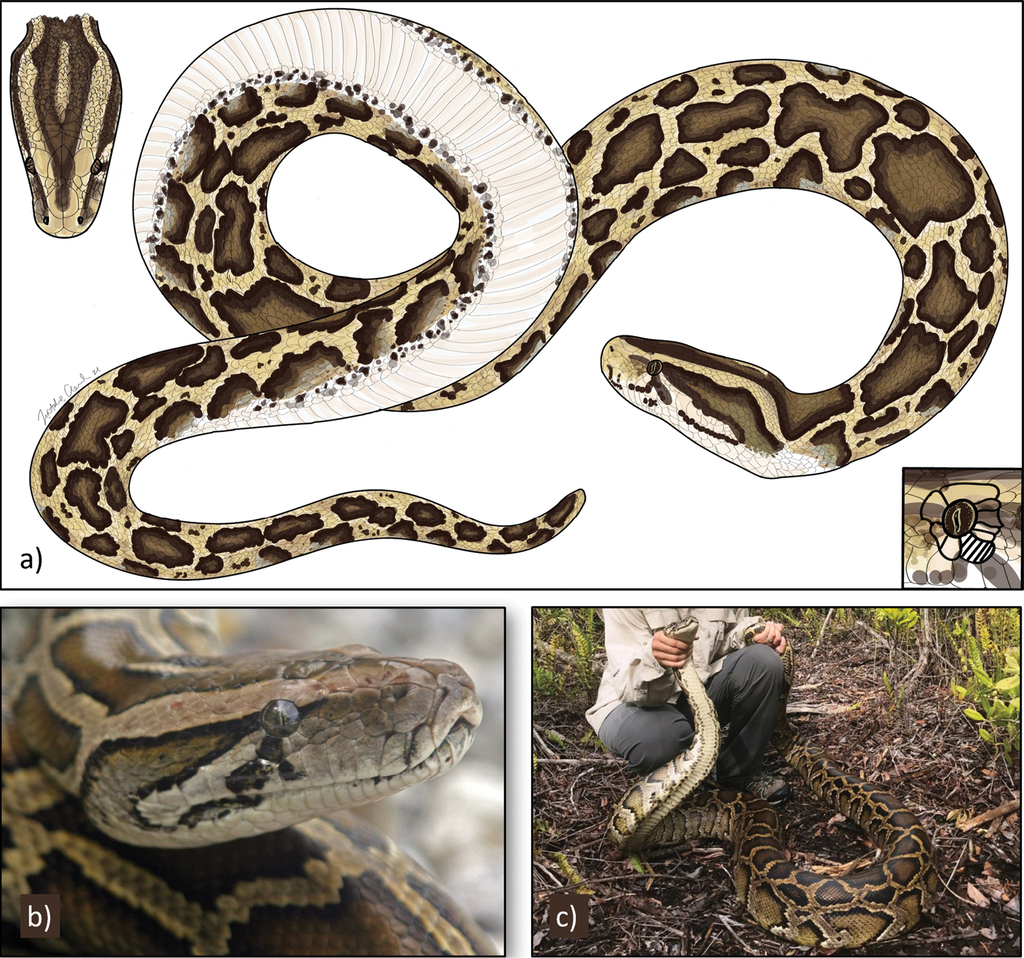
Burmese pythons have a distinct dorsal pattern of black-bordered, brown dorsal and lateral blotches separated by tan coloration that extends underneath to the venter (Fig.
Identification of Burmese pythons (Python molurus bivittatus). Distinguishing features include a, b a dark triangular spearhead on the top of the head extending to snout, with a white line extending posteriorly under the eye a subocular scale just below the eye preventing contact between supralabial scales and the eye (hatched lines, inset), and a, c a pattern of black-bordered, brown dorsal and lateral blotches separated by tan coloration and a white, non-patterned venter bordered by black spots. Panel a illustrated by Natalie Claunch. Photo credits: U.S. Geological Survey (b) and Conservancy of Southwest Florida (c).
Taxonomy
The status of the Burmese python as either a full species (Python bivittatus Kuhl, 1820) or a subspecies (Python molurus bivittatus Kuhl, 1820) of the Indian python (Python molurus Linnaeus, 1758) has been in flux for most of the past two centuries, and the taxonomy remains unstable today (
In general, there are geographic, morphological, and genetic characteristics suggesting that Indian and Burmese pythons may be distinct evolutionary lineages (i.e., they may fit the general lineage concept of species;
Taxonomic resolution for native-range Burmese pythons is of interest to the invasive population in Florida because there is evidence of hybridization between P. molurus bivittatus and P. molurus molurus - most likely before the introduction in Florida. In a sample of 426 pythons collected from locations throughout southern Florida between 2001–2012, two of six observed mitochondrial haplotypes were associated with 11 sequences from Indian pythons (
The geographic origin of ancestors to the Florida population is thought to be Thailand and Vietnam, based on the declared origin of most imports during the time when pythons were presumably introduced (
Ultimately, the large geographic range of Indian and Burmese pythons presents opportunities for isolation leading to speciation and undocumented cryptic diversity. Portions of the range are also thought to overlap, potentially allowing for historical mixing in those areas. Thus, more genetic information from the native range would be needed to resolve this issue (
Demography
Central to Burmese python management is understanding demography, or how vital rates (such as survival, growth, and reproduction) structure python populations. Changes in these birth and death processes drive changes in abundance over time and space (i.e., population ecology). Key demographic information can be summarized in a life table, which is a record of survival and reproductive rates in a population, broken down by age, size, or developmental stage (e.g., egg, hatchling, young of year, juvenile, sub-adult, adult). Information from life tables can be used to build a structured population model that can predict how changes to life-history parameters influence growth or decline of populations over time and thus how control tools might affect population dynamics. There is currently little information to construct a life table for Burmese pythons. However, future research to estimate life table parameters and develop a structured population model can help identify aspects of Burmese python life history of relevance to management efforts. For example, pinpointing the demographic parameters (e.g., age, stage, or size class) that contribute most to population growth can inform targeted removal efforts. An understanding of Burmese python vital rates can also identify potential hurdles to management efforts. For example, control efforts for invasive American bullfrogs (Lithobates catesbeianus) that focus on removing tadpoles and breeding adults can be offset by density-dependent competition and reduced cannibalism, so are less effective at decreasing population growth rate compared to seasonal culling of metamorphs (
Survival
Annual survival has not been well characterized for Burmese pythons, in part because requisite sample sizes and study durations for telemetry-based estimations are logistically and financially challenging (
As with adult pythons, there are few empirical data available to inform estimates of juvenile survival rates.
There are few published data on clutch or egg survival of wild Burmese pythons. However, brooding females are capable of shivering thermogenesis to raise embryonic temperatures during cool periods and exhibit parental care via nest attendance, which together likely increase embryo survival by discouraging potential nest predators (e.g.,
Mortality
In their native range, king cobras (Ophiophagus hannah,
Undocumented but potential predators of Burmese pythons in southern Florida include Florida panthers (Puma concolor coryi), coyotes (Canis latrans), foxes (Urocyon cinereoargenteus, Vulpes vulpes), various avian species, and invasive species like the Nile monitor (Varanus niloticus,
Humans are thought to be the primary cause of mortality for Burmese pythons in their native range, largely due to habitat loss and over-collection (
Reproduction
Burmese pythons in Florida typically mate over approximately 100 days during winter and early spring (early December to mid-March), when males seek and aggregate around mature females (
In southern Florida, females can have primary follicles throughout the year, develop secondary follicles most frequently from December into March, then oviductal eggs from March into May, and lay eggs in May (
Despite their prevalence in captivity (
In the low-elevation ecosystem of southern Florida, elevated habitats that remain relatively dry are important for nesting. In southwestern Florida where natural areas are interspersed with urban development, elevations of ~1.7 m have been associated with python nest site-selection, with nests concentrated on the urban fringe of the development, borders of agricultural fields, or in sandy upland habitat (
Clutch size of Burmese pythons increases with body size (
Current estimates of Burmese python (Python molurus bivittatus) demographic parameters by life stage in southern Florida, USA. Dash indicates data for parameter does not exist. Four example developmental stage classes are included. Asterisk (*) indicates Rookery Bay Estuarine Research Preserve and Collier-Seminole State Park. Reported clutch size values (^^) can include large pre-ovulatory follicles, oviductal eggs, or laid eggs, whereas data specific to oviductal eggs (i.e., those more likely to actually be laid) are indicated by a plus symbol (+) in the Notes column. Caret symbol (^) indicates data are based on individuals that have secondary follicles (i.e., pre-ovulatory, late-vitellogenic follicles).
| Annual survival | ||||||
|---|---|---|---|---|---|---|
| Age/Size Class | Estimate | 95% CI | Sample size | Location | Reference | Notes |
| Hatchling | 29% | 12–45% | 28 | southwest Florida* |
|
6-mo survival: 35.7% (21–60%); 2 of 28 survived 2yr post-release |
| Juvenile | – | – | – | – | – | – |
| Subadult | – | – | – | – | – | – |
| Adult | – | – | – | – | – | – |
| Fecundity | ||||||
| Reproductive frequency | Annual | Biannual | ||||
| – | – | 36% (n = 67 of 184) of adult ♀’s non-reproductive annually ( |
||||
| Clutch size | Python length | Clutch size^^ | Sample Size | |||
| SVL (cm) | TL (cm) | mean (range) | (# of clutches) | |||
| 264-286 | 297-322 | (21-37) | 7 |
|
||
| 184-292 | 210-328 | 22 (2-56) | 75 |
|
+ mean: 21 (2-41) | |
| 295-376 | 332-427 | 45 (27-74) | 27 | + mean: 39 (27-59) | ||
| 377-401 | 431-455 | 64 (42-86) | 11 | + mean: 60 (52-64) | ||
| 408-478 | 460-533 | 75 (35-103) | 16 | + mean: 53 (35-72) | ||
| 424-482 | 430-537 | (61-87) | 7 |
|
||
| Hatching rate | Number hatched | % Hatched | ||||
| 17 of 22 | 77 |
|
||||
| 50 of 61 | 82 |
|
||||
| 71 of 71 | 100 |
|
||||
| Minimum Female Size at Maturity | ||||||
| Minimum size at maturity | Python length | Sample size | ||||
| SVL (cm) | TL (cm) | |||||
| 185 | 210 | 2 |
|
|||
| Average minimum Size at maturity^ | 206♀, 182♂ | 80♀, 246♂ |
|
|||
| Age at maturity | – | – | ||||
| Longevity | – | – | ||||
Given brooding and nest-temperature maintenance by females, clutch survival is presumably high, and this is supported by data from a handful clutches, with hatching-success rates of 92% (n = 25 eggs;
Size distribution
Burmese pythons exhibit female-biased sexual size dimorphism with females larger than males, both in length, by as much as 150–180 cm, and mass, with the heaviest of females nearly twice as heavy as the largest males (85 kg vs 44 kg;
Burmese pythons in southern Florida vary in size from 34.44 to 500.9 cm SVL and 0.04–97.5 kg (n = 7,762; Figs
Distribution of Burmese python (Python molurus bivittatus) body size in southern Florida, USA (n = 9,501) varying from 34.44 cm to 500.9 cm SVL (snout-vent length). Bars represent pythons measured between 1995-2022 by state and federal agencies summarized by SVL. Data are from the U.S. Geological Survey and National Park Service (USGS/NPS, n = 3,723,
Relationship between snout-vent length (SVL; range: 34.44 to 500.9 cm) and mass (range 0.04 to 97.5 kg) of Burmese pythons (Python molurus bivittatus) measured in southern Florida, USA between 1995-2022 (n = 7,762). Data are from the U.S. Geological Survey and National Park Service (USGS/NPS, n = 3,723,
- Reproductive adult females:
- SVL (cm) = -4.25341 + 0.8954965*Total Length (cm)
- Total Length (cm) = 4.4790281 + 1.1165112*SVL (cm)
- Reproductive adult males:
- SVL (cm) = -3.287218 + 0.8818756*Total Length (cm)
- Total Length (cm) = 4.5850061 + 1.1287574*SVL (cm)
Python length-mass relationships are influenced by sex, where females are typically heavier per unit length than males, as well as length, where longer snakes are proportionally heavier per-unit length than shorter snakes (i.e., allometric growth;
Hatchling size
Burmese pythons are large snakes, even as hatchlings. In their native range, hatchling mass varies from 75–165 g and TL varies from 48–79 cm long (SVL ~43–71 cm;
Florida has 51 native snake species and Burmese python hatchlings are generally larger than neonates of the five largest native species (eastern diamond backed rattlesnake, Crotalus adamanteus: 30–38 cm SVL; ratsnake, Pantherophis obsoletus complex: 30 cm TL; eastern indigo snake: 45–61 cm TL; pine snake, Pituophis melanoleucus: 35–50 cm TL; coachwhip, Coluber flagellum: 30–44 cm TL;
Size at maturity
In the native range, females are considered mature at approximately 260 cm TL (~10 kg;
In the native range, the smallest reproductive male of the closely related Indian python measured 172 cm SVL (198 TL;
Maximum size
As with most ectotherms, maximum size is determined by factors such as environmental conditions, available resources, resource allocation decisions, and genetic variability (
Growth and longevity
Somatic growth patterns are a key life-history trait in all organisms and influence maximum body size which in turn affects survival, fecundity, and competitive ability (
Growth rates of wild pythons in Florida are not well-documented, in part because encountered individuals are generally removed rather than being marked, released, and recaptured to provide information on growth during the inter-capture interval. Captive feeding studies have documented Burmese pythons growing as fast as 20 cm per month (
In Florida, python age is only identifiable for the first several months of their first year, after which variation in individual growth makes it difficult to distinguish year classes (See Hatchling Size section). Pythons are approximately 100 cm SVL after one year (
Currently there are no longevity estimates for wild pythons, but the longest lifespans documented in captive Burmese pythons exceed 30 years (i.e., 28–34 yrs;
Physiology
Wild Burmese pythons can exhibit physiological resilience to stressors, including short-term captivity and handling (
Thermal biology
Burmese pythons are ectotherms that largely rely on environmental heat sources to control body temperature. However, they are active thermoregulators and use behaviors such as basking or seeking shade (
Body temperature influences behavior, physiology, and development (e.g.,
The aspect of thermal physiology most relevant to the Burmese python invasion in southern Florida is cold tolerance, with minimum temperature extremes being a stronger driver of survival than average temperature over a span of time (
To survive lethally cold air temperatures, pythons must retreat into sheltered refugia and remain there until temperatures warm again. Thus, understanding thermoregulatory behavior is critical to projecting range expansion beyond southern Florida. Burmese pythons in northern India appear to use refugia (e.g., porcupine burrows) to escape cold winter temperatures (
Osmoregulation
High salinity has been suggested as a limiting factor affecting reptile distribution in coastal habitats because very few species have adaptations (e.g., salt glands) to regulate salt uptake (
Hatchling Burmese pythons provided with marine water (salinity = 35 ppt) to drink survived in the lab for approximately one-month (mean = 32 days, 95% CI 23–40), whereas in brackish treatments (salinity = 10 ppt) pythons survived about five months (mean = 156 days, 95% CI 115–197 days;
Energetics and digestion
Python digestive physiology and energy budgets can be used to estimate rates of prey consumption, which are critical to understanding impacts on native species (see Diet and Prey Decline sections). Burmese pythons have unique morphological and physiological responses to feeding and fasting that likely have contributed to their success as invaders in Florida. Burmese pythons are thought to be primarily ambush predators (
An energy budget is a means of dividing ingested energy into allocations for metabolic processes and production (e.g., maintenance, growth, reproduction, fat storage) and for losses (i.e., in feces and urates;
There are no rigorous estimates of feeding frequency in wild Burmese pythons, because there are little data on pythons with empty stomachs. An examination of 1,716 pythons known to have contents in their digestive tract indicated the average number found in an individual python was 1.28 prey items, although one python contained 14 cotton rats (Sigmodon hispidus), including nine newborn rats (C. Romagosa, UF, Written Communication, 9/21/2022).
Detection probability
Broadly speaking, detection probability is the chance that an individual or species will be detected during a survey, given that it is present at the site. Snakes are generally considered the most difficult reptile group to study because of low detectability, resulting from secretive behavior, cryptic coloration, low and sporadic activity, and low abundance (
Examples of cryptic coloration contributing to low detection probability in representative habitat where Burmese pythons (Python molurus bivittatus) have been captured. White circles indicate pythons. Photographs illustrate python crypsis in hammocks (a–c), shrubs, mangroves, trees, (c, g, h) and cypress domes and wet prairies (d, e, f). Photo credits: Crocodile Lake National Wildlife Refuge (a, b, h) and U.S. Geological Survey (c–g).
Types of detection probability
There are two types of detection probabilities to consider depending on research or management goals: detection at the level of the ‘species’ or the ‘individual’. Species detection probability is used in occupancy studies and refers to the probability of directly or indirectly (e.g., from tracks or environmental DNA [eDNA]) encountering at least one individual of that species in a given area, given that at least one individual truly occupies the area. Individual detection probability, typically derived from capture-mark-recapture studies, refers to the probability of detecting a particular individual snake (
Species detection
The distinction between species and individual detection is important because species detectability reflects both abundance (i.e., how many individuals are present) and individual detection probability (i.e., how difficult each individual is to find). For example, species detection at a site may be considered high because an abundant species is detected on a large proportion of surveys, yet most individuals themselves will rarely be detected because of low individual detection probabilities. Understanding species detectability is critical for determining the geographic extent of an invasion, tracking spread, developing early-detection/rapid-response protocols, and assessing eradication status. Species detectability is generally used to determine whether a species is present at a given location and is frequently estimated at a population level using hierarchical models such as occupancy models (e.g., Pavlacky et al. 2012;
Environmental DNA as a detection tool for pythons
Environmental DNA is DNA released from an organism into the environment through sources such as feces, mucous, shed skin, hair, or decomposing carcasses (
Burmese python eDNA can be successfully amplified from water (
Because the Greater Everglades Ecosystem in Florida is a vast, shallow marsh with slow laminar sheet flow, eDNA monitoring has been efficient for species detection of Burmese pythons as compared to other tools such as visual searches, trapping, telemetry, or cameras (e.g., visual survey efforts resulted in <0.05 detection probabilities (
While eDNA is typically taken from water, it can also be amplified from soil samples for terrestrial detection of snakes. A laboratory experiment with captive bred corn snakes (Pantherophis guttatus) kept in terrestrial enclosures successfully identified eDNA in the soil for up to 96 hours post-removal (
Overall, eDNA is useful as a detection tool for cryptic species in that it does not rely on visual observations, utilizes readily available environmental samples and is not harmful to the environment or local species (
Challenges with eDNA
Although use of eDNA can minimize some aspects of labor-intensive traditional surveys (e.g., capture mark recapture, visual surveys) and thereby increase the ability to detect species, several challenges remain when using eDNA (reviewed by
Individual detection
For other management related questions (e.g., estimating abundance change over time), knowledge of individual detection probability is critical as it influences assessment of effective control strategies. A comprehensive understanding of individual detection probability has been a cornerstone of management initiatives for another invasive snake, the brown treesnake (Boiga irregularis) on the island of Guam (
Individual detection probability of Burmese pythons is extremely low. Using data from 59 visual surveys (144 person-surveys) for wild pythons along the C-110 canal near the Frog Pond Wildlife Management Area (Frog Pond WMA; Fig.
Low detection probability is a major challenge to python research and management because many surveys are needed to detect and remove most individuals within a population. In addition, estimation of demographic parameters such as abundance, survival, and recruitment (used to inform management) also generally depend on repeated detection of individuals over the course of many surveys that may span months or years (i.e., capture-mark-recapture;
Burmese python arrival in Florida
History of imports into the USA
Burmese pythons have been among the most heavily traded snake species for many decades (
Although the CITES import records are the best available metric of international trade, import records are unreliable (e.g., potential cross-border smuggling for subsequent re-export;
History of the invasive Florida population
Reports of escaped pet Burmese pythons in the United States have occurred since at least the early 1900s (
Geographic spread of Burmese python (Python molurus bivittatus) records in southern Florida between 1979 and 2021. Occurrence records were obtained from a large geospatial database of invasive species reports (Early Detection & Distribution Mapping System,
Between 2001 and 2003, many more Burmese pythons were documented throughout ENP, including from the Long Pine Key and Hole-in-the-Donut regions in the central part of the Park, the Chekika region on the eastern boundary of the Park, and the US-41, L-67, and Shark Valley regions in the northwest part of the Park (Figs
Introduction scenarios
Primary introduction
The initial source of Burmese pythons in southern Florida was the result of intentional or unintentional releases of captive pythons, and consideration of how they became established can be valuable for preventing establishment of similar species in the future.
Possible secondary introduction
There are several lines of evidence suggesting a possible second introduction of Burmese pythons to southwestern Florida. Burmese pythons were seen in the area outside of Naples and along US-41 by credible observers beginning in the late 80s and into the 90s (I. Bartoszek, CSWFL, Written Communication, 5/27/2021). The first Burmese python records in southwestern Florida occurred in 2003 and 2005, in western Collier County, near Everglades City, approximately 5 and 22 km east, respectively, of the intersection of US-41 and Hwy-29 (Fig.
Population genetics
The earliest population genetics work examined 156 individual wild Burmese pythons collected in ENP between 2003–2006 and compared them to a single shed skin from a local pet store and 13 skins from a local reptile dealer that were purportedly from a wild population in Vietnam (
Status of the Florida population
Abundance
Estimating abundance and population growth rates of wild python populations is vital for effective management, but it is difficult to obtain accurate estimates, particularly for cryptic invasive species. Although unsubstantiated ranges of abundance are frequently circulated in the popular media, there are no reliable estimates of python abundance or density. This knowledge gap makes it challenging to evaluate effectiveness of current or proposed control methods and management initiatives. A common method for abundance estimation is using capture-mark-recapture (CMR) surveys where many individuals are marked and released, and the proportion of marked individuals recaptured later can provide robust estimates of population size (e.g.,
Challenges interpreting removal data
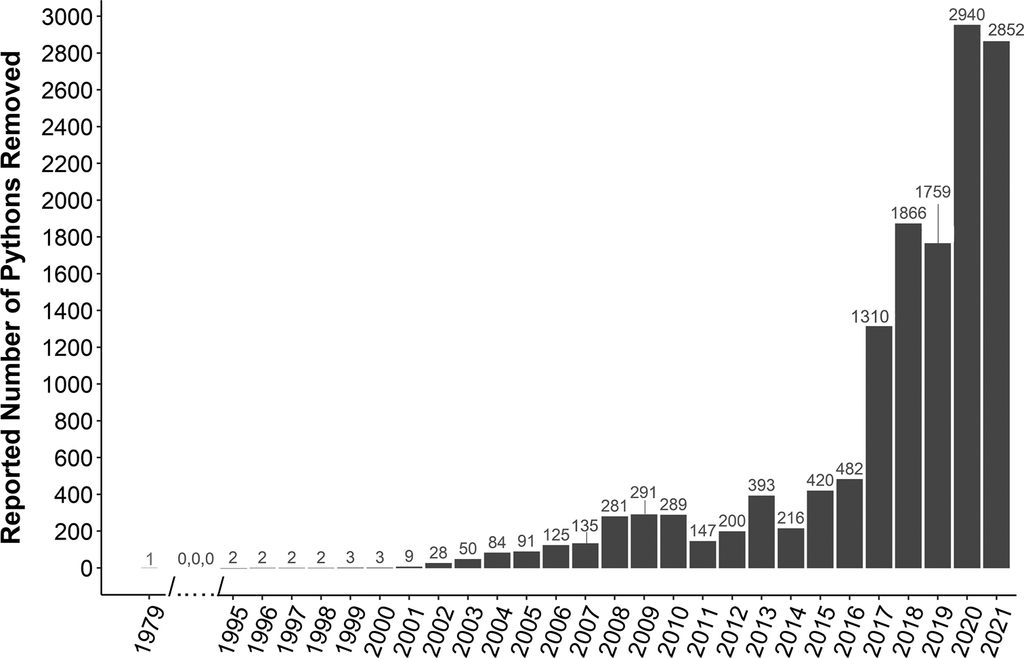
Although less rigorous, annual removal data (raw counts) can provide an approximate index of relative abundance for given areas.
Annual number of reported removals of Burmese pythons (Python molurus bivittatus) across southern Florida through December 31st, 2021 (n=13,746). Data reported to and managed by Florida Fish and Wildlife Conservation Commission (FWC; Suppl. material
Records from ENP are consistent with a population decline and subsequent growth following an extreme cold event in January 2010. Python removals from Main Park Road in ENP from 2000–2014 increased dramatically from less than 10 per year in 2000 and 2001, to a peak of ~140 per year in 2009 (
An important component of Burmese python ecology is an understanding of the interactive effects of removals and natural mortality (
Density
There is little to no information on python abundance across southern Florida. Abundance is typically estimated using capture-mark-recapture studies of individuals, but because individual detection rates in pythons are extremely low, encounter rates are extremely low and re-encounter rates are even lower, making the cost and duration of this kind of study of pythons an obstacle (see Detection Probability section). Agricultural activities have allowed for rough density estimation at Frog Pond WMA on the eastern edge of ENP (505 ha total area, Fig.
The only attempt to rigorously estimate python density in southern Florida is a novel, simulation‐based technique developed by
Several factors may have biased
Density estimates of 1.5 to 5 pythons per km2 in ENP may be low, especially considering that large, native snake species often exist at densities greater than one per ha (100 per km2;
Overall, there is little information about python population size within ENP, and no information regarding abundance across southern Florida. The standard technique for robust abundance estimation while accounting for imperfect detection of all individuals in the population is capture-mark-recapture (e.g.,
Geographic distribution
The most common method for determining distribution of any species is to plot occurrence records. In the case of Burmese pythons, this approach can depict the spatial spread of snakes over time (Fig.
Range in Florida
Determining the python distribution is an ongoing research priority but is challenging because the invasion front is shifting over space and time, much of the landscape is inaccessible, survey effort is not available for many records, and python detection probability is extremely low (see Detection section). However, there is a large geospatial database of invasive species reports, Early Detection & Distribution Mapping System (EDDMapS; https://www.eddmaps.org), that includes Burmese python observations submitted by python researchers, land managers, and the public. Though distribution records are biased towards accessible areas (levees and roads; Fig.
Based on available records, Burmese pythons occupy most of southern Florida, encompassing approximately 30,000 km2 from Lake Okeechobee throughout Palm Beach County, south through Miami-Dade County to Key Largo, and west throughout Monroe, Collier, Hendry, and Lee Counties (Fig.
Early detection using eDNA
By 2013, the most northern samples documenting Burmese python eDNA were taken south of Lake Okeechobee in Holey Land Wildlife Management Area and the adjacent Stormwater Treatment Area (STA) 5 (Fig.
Potential range
To project the potential range of Burmese pythons in the United States, several species distribution models have been produced with the goal of characterizing the climate of their native range in southeast Asia and identifying sites elsewhere in the world that may be climatically similar and therefore at risk of invasion by pythons. These endeavors provoked public controversy (reviewed in
Overall, potential range limits of Burmese pythons are uncertain. Multiple climate matching efforts have reached different conclusions. In addition, there is evidence that evolutionary change has already altered parts of the genome responsible for cold tolerance (see Cold Tolerance section), and there is the potential for behavioral plasticity to enhance cold tolerance by pythons seeking refugia (see Refugia section). Further, climate change is ongoing and may be outpacing previous climate projections (
Refugia
Species distribution models have overlaid the range of climate conditions from occupied areas of the native range onto the United States; however, pythons may be able to occupy an expanded climate envelope if released from native-range biotic pressures (e.g., prey availability, competition, predation, refugia, and disease) and if available refugia exist. For example, Burmese pythons use gopher tortoise and mammal burrows as refugia (
Cold tolerance
Movement
Movement ecology is an ecological subdiscipline that connects an animal’s movement path with environmental heterogeneity, available resources, motion and navigation capacity, and its internal state (e.g., motivation;
Adult Burmese pythons are capable of long-distance movements of several km (
In southern Florida, an extensive network of canals and levees may facilitate long-distance movement by pythons, and consequently, expansion across the landscape. However, movement in canals may be a risky strategy due to predation by alligators (
Navigation and homing
Navigational ability is the process by which animals decide when and where to move (
Dispersal
Dispersal is the movement of organisms away from their place of birth (
Home range
An animal’s home range is most commonly defined as the area it uses during the course of normal activities such as foraging and mating but excluding occasional exploratory excursions (
Habitat use
Aside from potentially differing prey availability, the varying habitats in Florida may also influence movement and home range of pythons. For example, although both ENP and southwestern Florida contain extensive wetland habitats, within southwestern Florida these wetlands are interspersed throughout xeric, upland habitat types that are more predominant than within the “river of grass” ridge and slough system that characterizes ENP (
Another potentially important factor that may influence home range in southern Florida is urban landcover, which is permeated by an extensive network of canals and drainage ditches that facilitate movement by pythons (
Most research on habitat selection has been at the larger scales of home range and use of habitat types within, but factors characterizing nest site selection in Florida beyond drier elevated habitats are poorly understood. Several single observations have been described in which females selected highly modified habitats such as debris piles (
Burmese python impacts
Human safety
There are no reports of humans being killed by wild Burmese pythons in Florida; instead, the few recorded deaths have occurred from captive pythons. People have been bitten by wild juvenile and adult Burmese pythons in Florida, but these events are typically provoked during capture attempts (
Overall, there appears to be very low risk of unprovoked, serious human injury or fatality by wild Burmese pythons in southern Florida (
While there is little risk of python-induced injury to humans, there may be some threat from consumption of python flesh. Mercury concentrations are high in tissues of pythons collected from southeast Florida (mean 4.35 mg/kg, n = 136;
Although direct risks to humans may be minimal, python-induced changes to mammal community composition (
Direct ecological impacts on wildlife
Burmese pythons consume a wide range of vertebrate prey, particularly mammals, and directly influence and alter food webs throughout southern Florida. Invasive species, particularly invasive mammalian predators, have contributed to extensive global species declines and extinctions (
Foraging strategy
Burmese pythons are considered ambush predators that eat infrequently but consume a wide variety of terrestrial vertebrate prey (
Despite the general acceptance that pythons are ambush predators, there are few observations in the wild of this behavior.
Diet
In both their native range and in southern Florida, Burmese pythons are dietary generalists that consume a wide range of vertebrate prey, consisting mostly of mammals and birds, although there are a few records of lizards, frogs, and snakes from the native range (
Prey species found within digestive tracts of Burmese pythons (Python molurus bivittatus) in southern Florida. Threat status under the United States Endangered Species Act (Gruver and Montero 2018) is indicated where applicable in bold: FE, federally endangered; FT, federally threatened; FT-S/A, federally threatened due to similarity of appearance, ST, state threatened. Reprinted from (
| Order | Family | Scientific Name | Common Name | Count |
|---|---|---|---|---|
| Aves | ||||
| Accipitriformes | Cathartidae | Coragyps atratus | Black Vulture | 7 |
| Anseriformes | Anatidae | Aix sponsa | Wood Duck | 2 |
| Anas acuta | Northern Pintail | 1 | ||
| Anas crecca | Green-Winged Teal | 1 | ||
| Anas domesticus | Domestic Mallard | 1 | ||
| Anas fulvigula | Mottled Duck | 1 | ||
| Anas platyrhynchos | Mallard | 2 | ||
| Spatula discors | Blue-Winged Teal | 2 | ||
| Charadriiformes | Scolopacidae | Gallinago delicata | Wilson’s Snipe | 2 |
| Numenius phaeopus | Whimbrel | 1 | ||
| Ciconiiformes | Ciconiidae | Mycteria americana (FT) | Wood Stork | 3 |
| Columbiformes | Columbidae | Zenaida macroura | Mourning Dove | 1 |
| Galliformes | Phasianidae | Gallus gallus | Chicken | 5 |
| Numida meleagris domesticus | Guineafowl | 2 | ||
| Gruiformes | Aramidae | Aramus guarauna | Limpkin | 21 |
| Rallidae | Fulica americana | American Coot | 20 | |
| Gallinula galeata | Common Gallinule | 11 | ||
| Porphyrio martinica | Purple Gallinule | 4 | ||
| Porzana carolina | Sora | 8 | ||
| Rallus elegans | King Rail | 17 | ||
| Rallus limicola | Virginia Rail | 1 | ||
| Rallus longirostris | Clapper Rail | 2 | ||
| Passeriformes | Cardinalidae | Cardinalis cardinalis | Northern Cardinal | 2 |
| Corvidae | Corvus brachyrhynchos | American Crow | 1 | |
| Icteridae | Agelaius phoeniceus | Red-Winged Blackbird | 3 | |
| Dolichonyx oryzivorus | Bobolink | 1 | ||
| Quiscalus major | Boat-Tailed Grackle | 2 | ||
| Sturnella magna | Eastern Meadowlark | 3 | ||
| Mimidae | Dumetella carolinensis | Gray Catbird | 1 | |
| Mimus polyglottos | Mockingbird | 1 | ||
| Parulidae | Geothlypis trichas | Common Yellowthroat | 1 | |
| Pycnonotidae | Pycnonotus jocosus | Red-Whiskered Bulbul | 1 | |
| Regulidae | Regulus calendula | Ruby-Crowned Kinglet | 1 | |
| Troglodytidae | Troglodytes aedon | House Wren | 3 | |
| Pelecaniformes | Ardeidae | Ardea alba | Great Egret | 7 |
| Ardea herodias | Great Blue Heron | 6 | ||
| Botaurus lentiginosus | American Bittern | 19 | ||
| Butorides virescens | Green Heron | 4 | ||
| Egretta caerulea (ST) | Little Blue Heron | 2 | ||
| Egretta thula | Snowy Egret | 2 | ||
| Ixobrychus exilis | Least Bittern | 3 | ||
| Nycticorax nycticorax | Black-Crowned Night-Heron | 4 | ||
| Threskiornithidae | Eudocimus albus | White Ibis | 44 | |
| Platalea ajaja (ST) | Roseate Spoonbill | 1 | ||
| Podicipediformes | Podicipedidae | Podilymbus podiceps | Pied-Billed Grebe | 41 |
| Suliformes | Anhingidae | Anhinga anhinga | Anhinga | 6 |
| Fregatidae | Fregata magnificens | Magnificent Frigatebird | 2 | |
| Suliformes | Phalacrocoracidae | Phalacrocorax auritus | Double-Crested Cormorant | 1 |
| Mammalia | ||||
| Artiodactyla | Bovidae | Capra aegagrus hircus | Domestic Goat | 1 |
| Cervidae | Odocoileus virginianus | White-Tailed Deer | 43 | |
| Suidae | Sus scrofa | Wild Boar | 3 | |
| Carnivora | Canidae | Urocyon cinereoargenteus | Gray Fox | 4 |
| Felidae | Felis catus | Domestic Cat | 5 | |
| Lynx rufus | Bobcat | 5 | ||
| Mustelidae | Lontra canadensis | River Otter | 1 | |
| Procyonidae | Procyon lotor | Raccoon | 43 | |
| Cingulata | Dasypodidae | Dasypus novemcinctus | Nine-Banded Armadillo | 3 |
| Didelphimorphia | Didelphidae | Didelphis virginiana | Virginia Opossum | 77 |
| Eulipotyphla | Soricidae | Blarina carolinensis | Southern Short-Tailed Shrew | 3 |
| Cryptotis parva | North American Least Shrew | 3 | ||
| Lagomorpha | Leporidae | Sylvilagus floridanus | Eastern Cottontail | 39 |
| Sylvilagus palustris | Marsh Rabbit | 52 | ||
| Rodentia | Cricetidae | Neofiber alleni | Round-Tailed Muskrat | 94 |
| Neotoma floridana | Eastern Woodrat | 13 | ||
| Neotoma floridana smalli (FE) | Key Largo Woodrat | 10 | ||
| Oryzomys palustris | Marsh Rice Rat | 28 | ||
| Peromyscus gossypinus | Cotton Mouse | 120 | ||
| Peromyscus gossypinus allapaticola (FE) | Key Largo Cotton Mouse | 3 | ||
| Sigmodon hispidus | Hispid Cotton Rat | 528 | ||
| Muridae | Mus musculus | House Mouse | 20 | |
| Rattus norvegicus | Brown Rat | 1 | ||
| Rattus rattus | Black Rat | 333 | ||
| Sciuridae | Sciurus carolinensis | Gray Squirrel | 3 | |
| Sciurus niger | Fox Squirrel | 4 | ||
| Reptilia | ||||
| Crocodilia | Alligatoridae | Alligator mississippiensis (FT-S/A) | American Alligator | 71 |
| Squamata | Iguanidae | Iguana iguana | Green Iguana | 1 |
| Total number of species | 76 | |||
| Total number of diet items | 1788 | |||
Several methods have been used to identify prey from within Burmese python gastrointestinal tracts including morphological, molecular, and isotopic techniques. Thus far, morphological methods including microscopy have been the most successful for individual prey identification (
Since early 2000, several organizations including the United States federal government (U.S. National Park Service, U.S. Geological Survey), state of Florida (Florida Fish and Wildlife Conservation Commission, South Florida Water Management District), universities (University of Florida), and nonprofits (Conservancy of Southwest Florida) have been removing Burmese pythons across southern Florida and conducting necropsies to identify gastrointestinal contents for a direct account of diet in their invaded range. Expanding on prior studies,
Number of prey by order (y-axis) and family (text within bars) found within stomachs of Burmese pythons (Python molurus bivittatus). Black silhouettes are a general representation of animals within each family. Asterisks indicates several unlabeled families including eight within Passeriformes (Icteridae, Troglodytidae, Cardinalidae, Corvidae, Mimidae, Parulidae, Pycnonotidae, Regulidae), three within Suliformes (Anhingidae, Fregatidae, Phalacrocoracidae), and two within Artiodactyla (Suidae, Bovidae). Data from Table
While mammals were more numerous within python gastrointestinal tracts, birds made up the most diversity. Of the 48 species or subspecies of birds identified in python guts, wading birds (Pelecaniformes, n = 10 species, 20.8%), songbirds (Passeriformes, n = 12 species, 25%), and rails (Rallidae, n = 7 species, 14.6%) comprised most of the bird species diversity (Table
Species of concern
The presence of large species such as bobcat, deer, as well as a wide variety of highly mobile bird species in python gastrointestinal tracts indicates that almost any native endotherm within southern Florida is vulnerable to predation by pythons. Several species of concern have been documented in python stomachs, including state-threatened species [little blue heron (Egretta caerulea), roseate spoonbill (Platalea ajaja), Big Cypress fox squirrel (Sciurus niger avicennia)], federally threatened species [wood stork], and federally endangered species [Key Largo woodrat (Neotoma floridana smalli;
In addition to direct predation, Burmese pythons may compete with native species such as bobcats, Florida panthers, predatory birds, and snakes (e.g., eastern indigo and eastern diamondback) for prey. These native predators also have broad diets and consume a variety of birds, small and mid-sized mammals, and reptiles. In the case of panthers, competition may occur for large mammalian prey such as white-tailed deer and wild hogs (
Ground-dwelling birds such as cranes, coots, and gallinules, as well as wading birds (ibises, storks, spoonbills, egrets, herons) may be particularly at risk because they lack the reproductive output typical of mammals such as rodents or feral hogs, and all life stages (eggs, young, adult) are susceptible to predation by native carnivores, as well as pythons (
Mammal declines
Burmese pythons in southern Florida consume a wide range of mammals (
Subsequently,
Threats to the Greater Everglades Ecosystem not only include invasion of non-native species but also alteration of hydrology throughout the system, water quality deterioration (e.g., nutrients, sulfate, mercury, pesticides, heavy metals), agricultural and urban development, altered disturbance regimes, and climate change (
Indirect ecological impacts
There is considerable research focusing on direct negative impacts of Burmese pythons in southern Florida. However, indirect effects may profoundly affect native ecosystems, including the spread of pathogens (e.g., serpentovirus) and parasites to native species, alteration of host-parasite dynamics (e.g.,
Parasites and pathogens
In their native range, Burmese pythons are host to ectoparasites (e.g., ticks, mites), blood parasites (e.g., hemogregarines), protozoan intestinal parasites (e.g., coccidia), metazoan intestinal parasites (e.g., tapeworms, roundworms), and other endoparasites (e.g., pentastomes;
In southern Florida, several parasite species infect pythons, including native North American snake parasites (e.g., pentastome, Porocephalus crotali,
About 13% of pythons in Florida are infected with R. orientalis (
Given the diverse assemblage of available hosts in North America, R. orientalis may expand both to new areas and new host species (
In addition to internal parasites, host switching among non-native and native external parasites have been documented in pythons. Ectoparasites on Burmese pythons have been documented in southern Florida, including non-native ticks (Amblyomma rotundatum and A. dissimile) and two species of chiggers (Eutrombicula splendens and E. cinnabaris) native to the United States (
Burmese pythons have been documented carrying a snake-associated virus in the order Nidovirales that causes respiratory disease (i.e., serpentovirus, reviewed by
Trophic-structure changes
Brown treesnakes are widely recognized as having caused ecosystem effects on Guam by extirpating or suppressing most of the island’s vertebrate species, including important pollinators and seed dispersers, thereby influencing forest diversity (
Declines in mesomammal predators may also indirectly affect other trophic levels, given that many are well-known as predators of reptile nests, particularly turtle nests (e.g., raccoons, opossums, and foxes;
Changes in food web structure or ecosystem services are often intricate and difficult to study and predict, especially in complex food webs where the possibility for compensation or redundancy of species’ roles may exist (
Python-induced mammal declines have also indirectly influenced transmission of zoonotic pathogens. Everglades virus is a mosquito-borne (C. cedecei) zoonosis of the Venezuelan equine encephalitis complex that is endemic to Florida and causes occasional nonfatal neurological disease in humans (
What do we know about control tools?
Management of invasive reptiles is a complex and challenging problem. A wide range of tools and techniques are available to detect and capture snakes (
Methods used to detect and capture Burmese pythons (Python molurus bivittatus). Abbreviations for the timings of application include year-round (YR), breeding season defined as December through March (BS), and late summer (LS) during the hatchling dispersal window which occurs in August. Abbreviations for life stage targeted include reproductive adult (RA), all size classes (All) and all size classes, but dependent on prey size (All*). The cost estimates may vary according to management area or agency and may change over time.
| Methods to locate and/or capture Burmese pythons | Timing of Application | Life Stage Targeted | Primary Use | Key Limitations | Cost Estimate | References |
|---|---|---|---|---|---|---|
| Visual Surveys | ||||||
| Visual Surveys: road cruising | YR, LS | All | Removal | Most of the landscape is > 1 km from a road and these areas can sustain populations | ~$298 per python removed (python contractors) |
|
| Visual Surveys: diurnal pedestrian | BS | All | Removal, particularly reproductive adults | Inefficient and costly because of low individual detection (i.e., <0.05) |
|
|
| Scent detection dogs | YR | All | Detection, Rapid Response, Range Delimitation | Hot/humid temperatures and dense vegetation limit dog performance | Can be as low as $65-100k/yr; $150-250/km; $730-$1,520/day |
|
| Artificial refugia | YR | All | Detection, Removal | Low yield method to detect pythons |
|
|
| Burrow camera | YR | All | Detection | Low yield method to detect pythons |
|
|
| Tracking | ||||||
| Scout snakes | BS | RA | Removal, particularly breeding females | Labor intensive, expensive | ~$11k/python; less expensive with less air support Some programs as low as $1,800/python |
|
| Telemetry (of prey animals) | YR | All* | Detection, Removal | Inefficient, low yield |
|
|
| Trapping | ||||||
| Baited trap (python prey, female python) | YR | All | Detection, Removal; Indirectly survey until trap is sprung | Lack of suitable attractant or traps that pythons will enter | Traps less expensive (<$200/trap); labor and vehicle cost are more expensive |
|
| Drift fence | YR | All | Detection, guide python into a trap | Height and material to prevent climbing over; thus far pythons do not readily enter traps | ||
| Burrow trap | YR | All | Removal | Requires previous knowledge of python presence in burrow, frequent monitoring |
|
|
| Camera traps | YR | All | Detection, Indirect and continuous surveying | Potentially millions of photos can result; automated algorithms cannot yet identify pythons; Does not result in capture |
|
|
| Biological | ||||||
| eDNA | YR | All | Detection, Range Delimitation | Does not result in capture; DNA can be transported |
|
|
| Pheromones | BS | RA | Removal | Pheromonal lures have not yet been well-developed or successful | ||
| Mechanical | ||||||
| Infrared (handheld gun/drone) | YR | All | Detection, Removal | Infrared technology is still being explored as a tool to increase detections during road cruising |
|
|
| Mowing/discing | YR, BS | All | Detection, Removal | Limited to agricultural lands and easements along levees |
|
|
Visual and road surveys
Burmese pythons are rarely seen, and even pythons located in accessible areas (i.e., adjacent to roads and levees) or outfitted with radiotransmitters are very difficult to find because they are usually concealed in vegetation or under water (
Removal programs
The SFWMD, FWC, and NPS have implemented python removal programs to pay contractors or authorize trained volunteers to capture and remove pythons across southern Florida (
Cost of visual surveys
Over the course of two years from 1 May 2019 to 30 April 2021, FWC contractors (see Removal Programs) conducted 4,731 surveys and removed 2,107 Burmese pythons (
Future applications of visual surveys
Although visual surveys currently result in the highest number of Burmese python captures (see Removal Programs section), it is unclear whether intensive road removals can act as resource protection (i.e., providing localized suppression to protect road-adjacent prey communities over the long term). While visual surveys are currently the most-used control tool for Burmese pythons, the vast roadless areas across southern Florida offer enough suitable habitat to sustain the python population indefinitely and serve as a source for recolonization of the roadside areas (see Challenges Interpreting Removal Data section). Thus, a combination of visual surveys and several other detection and control methods would likely be necessary to suppress the population of Burmese pythons across the entire occupied range, if it is possible at all. In addition, advances in technology may reduce cost per python removed. For example, human eyesight can detect light with wavelengths from 400 to 700 nanometers (nm;
Scout snakes
The scout technique uses radiotelemetry to capitalize on social behaviors of animals (e.g., seasonal aggregation) to improve detection and to reduce nuisance or invasive populations (
Although pythons are not typically social, from December to March in Florida they may form breeding aggregations that have been observed to include up to eight pythons (
To evaluate the scout snake technique,
In areas where multiple control tools are deployed simultaneously (see Removal Programs section), there is a risk of accidental removal of scout snakes, but this can be mitigated by external markers and effective communication. Thus far, the primary method used to externally mark scout pythons in southern Florida is with brightly colored polyolefin tubing (T-bar style Floy tags) shrink-wrapped around monofilament tags and anchored subcutaneously (e.g.,
Cost of scout snakes
Using data from five seasons of scout snake radiotelemetry efforts,
In general, the costs of using scout snakes are expected to vary widely, depending on location, habitat accessibility, and aviation time and type; programs that do not rely heavily on flight support are likely substantially less expensive per python removed. Personnel in fixed-wing aircraft (i.e., small airplanes) can find the general locations of telemetered snakes and guide technicians on the ground, and helicopters may be necessary to transport staff to and from remote sites where other means of transport are not viable (
Future applications of scout snakes
Although scout snake programs are more costly per python removed than road cruising, scout snakes are a tool targeting the removal of large, reproductive pythons that are far from roads and that might not be captured otherwise (see Detection section). This technique is implemented during the breeding season when pythons aggregate, and thus scout snake programs may reduce overall costs by increasing tracking frequency during the breeding season and reducing tracking during the remainder of the year. Although both male and female Burmese pythons lead researchers to breeding aggregations and are similarly effective in that they result in similar numbers of pythons removed, programs that use male pythons as scout snakes may lead to the removal of more females (
Because there may be individual variation in reproductive activity each breeding season among pythons, there is not yet a consensus on what traits make a scout python successful at finding other pythons. Scout success may vary by python density and is complicated by search difficulty in some habitats, but unlike removals by human searchers, the technique is not limited to roads and levees. Ultimately, very high, sustained funding would be required to determine if the scout snake technique can be scaled to a level of effort large enough to impact the population. Important considerations for using scout pythons are their powerful homing and navigational abilities (see Navigation and Homing section;
Overall, scout program costs do not scale linearly with the number of scout snakes, and there may be opportunities to leverage economies of scale with a larger scout program. While all scout snakes provide important life history data (which are currently limited; see Demography section), the criteria that define a productive scout snake vary according to program objectives. Some scout snakes may make large movements in and out of focal areas, rendering them more difficult to consistently track, whereas others may less consistently locate other pythons; in both cases those scouts increase program costs. Similarly, male scouts may locate females that a research program may opt to leave in the wild and track to obtain demography data; in these cases, male scouts may stay with that female, thus reducing opportunities to locate additional pythons for removal. Where management is the primary goal and in cases where particular scouts appear to be underperforming, it may be necessary to eliminate them to increase efficiency of removals.
Trapping
Using traps to catch snakes circumvents the need for an observer to locate individuals, reducing bias and making it possible to survey difficult habitats during all hours or until a trap is sprung. For decades, snakes have been successfully captured by traps equipped with funnel-style entrances (
Traps deployed in southern Florida that are sized for pythons need to provide avenues for escape or release of non-target species. Among the key non-targets are endangered or threatened species such as Key Largo woodrat and Eastern indigo snake, as well as species typically destructive to traps (e.g., raccoons, alligators, rats), sensitive species (e.g., birds), and venomous snakes because they are dangerous to remove from traps (
Experiments
Funnel trapping
To assess the efficacy of traps for Burmese python population control,
Large reptile traps
Preliminary trials in outdoor enclosures examined effectiveness of the large reptile trap (i.e., a Tomahawk model 463) with three large native snakes (2 cottonmouths and 1 yellow rat snake, Pantherophis alleghaniensis); results were promising, as native snakes did not trigger the traps to close, but pythons did (
Cost of trapping
Trap costs vary by design but a large reptile trap (i.e., Tomahawk Model 463) with a shade cover costs approximately $185. Thus far, large reptile traps used in
Challenges with trapping
Operational costs can restrict large-scale trapping efforts (reviewed in
Thus far, the main practical failing of traps appears to be a lack of either a suitable attractant or traps that pythons will enter (see Foraging Strategy and Trapping Experiment sections). Instead of prey bait, other attractants may prove useful for luring and capturing pythons in traps, including sex pheromones (see Pheromone section). However, pheromonal lures for use in traps have not been developed, and it is unclear how aspects of pheromone volatility and distance may influence their effectiveness. Other trap efforts may include large, portable (i.e., lightweight, collapsible) traps deployed during the breeding season, containing a wild-caught reproductive female as an attractant; trials with these traps are ongoing but thus far have not been effective (M. McCollister, NPS, Written Communication, 9/19/2022). In conjunction with different attractants, captive and field experiments with traps could incorporate drift fences along with replicated trap trials in python-occupied natural habitats with varying resource densities (e.g., mates, prey). Drift fence arrays are designed to intercept moving animals (
Overall, efforts thus far suggest that trapping for Burmese pythons is not currently a viable method of population control or eradication because python movement and behavior renders the snakes unlikely to regularly encounter traps. Ultimately, traps are one of a variety of control tools that managers may choose from. Future modifications that result in consistent python captures might make traps a cost-effective tool for local python control, or aid in early-detection efforts in newly invaded areas (
Pheromones
Scent trailing using pheromones is the principal mode of reproductive communication in snakes, allowing individuals to locate each other by detecting chemical compounds from the skin (
Scent detection dogs
The powerful olfactory receptors of dogs have made them useful for locating several groups of invasive, cryptic, or rare species, including plants, tortoises, birds, mammals, and snakes (reviewed in
Dog characteristics important for python detection include a strong play drive, independence, and confidence because they must work for long periods of time. They must also be in very good physical condition given the temperatures, terrain, and long distances required for searching (reviewed in
Overall, dogs can be used as a complement to other control tools. Dogs are most useful in situations where chance of human detection is low, such as assessment of python presence along areas peripheral to their current spatial range (e.g., Fig.
Cost of scent detection dogs
The cost estimate for detection dogs (as with all control tools) may vary according to management area or agency and may change over time. The purchase and maintenance of a dog program from an established canine performance program over the 8-year work life span of two dogs was estimated in 2011 to be approximately $561,200 (first year: $106,200, following 7 years: $65,000 /year;
Toxicants
Several commercially available products are lethal to snakes when ingested (
Control tool summary
Existing control tools outlined thus far may work well in combination, and on a small scale within a narrow timeframe or range of circumstances (Table
Future research
Over the past two decades, we have learned much about the biology and management of Burmese pythons. However, most of this information comes from isolated, relatively small-scale studies at few locations, or extensive incidental (i.e., not question-driven) data derived from python removals across southern Florida. Recently, multiple federal, state, and non-profit entities have combined efforts to accomplish targeted long-term studies at broader scales. This approach will attempt to generate estimates of abundance, detection probability, and vital rates for effective decision making and management of the python population (see Demography section). Suppressing Burmese python populations throughout vast and complex wilderness habitats that are managed by many different government, state, tribal, and private entities is a daunting management challenge that can be strengthened by basic and applied research studies. Below we outline some general research strategies and themes for future work.
Population suppression
Burmese pythons are now established across a large area of southern Florida, minimally encompassing areas from Palm Beach County, south to Key Largo, and west throughout Collier County (Figs
Refinement of existing control tools
Over the past two decades, much effort has gone into exploring control tools for Burmese pythons. Existing technologies (road surveys, visual surveys, scout snakes) that can result in python captures have advanced from initial research ideas to implementation by management agencies. Nonetheless, there are ample opportunities to refine these control tools, and some of these opportunities are described above (see Control Tools section).
Relatively little research has rigorously attempted to quantify detection via these methods (but see
Baseline abundance estimation
Overall, despite removal of many Burmese pythons over the past several decades (Fig.
For removal models to be informative they must (1) account for effort, (2) be targeted to a defined area, ideally with effort equally spread across the area to minimize heterogeneity in detection, and (3) have enough removal pressure to result in a substantial reduction in abundance. For example,
Removal approaches may also provide additional information to evaluate management actions by incorporating close-kin mark-recapture to infer population demographics. Close-kin mark-recapture identifies close-kin pairs (e.g., parent-offspring, half siblings) using genetic sampling of individuals and has been applied to assess relatedness of fish species in both fresh and saltwater systems to infer population sizes (
Relative abundance and abundance indices
Once population abundance has been estimated for a given location, a baseline exists to monitor resulting shifts in abundance. After developing a baseline, simpler and less-expensive methods such as an index of abundance could be evaluated (and calibrated) to track changes in abundance over time and space (
Expanding current removal efforts (e.g., see Removal Programs section) may provide an avenue to track changes in relative abundance over time. For example, a useful initial index to compare with more rigorous abundance estimates could include systematic road surveys (without removal) at regular locations over time, while recording number of pythons along with search effort and variables expected to influence python detection (e.g., weather, survey hour, observers, season). Developing initial abundance indices using road surveys in focal areas or research sites would require coordination among agencies conducting removal efforts to ensure standardized surveys where pythons were recorded but not removed. Improving current removal protocols to include surveys within a larger grid area overlapping road transects, where researchers would survey throughout the area using a mark-recapture framework with radiotagged pythons (see Abundance section), may provide information for abundance as well as survival estimation (e.g., known fate analysis;
Long-term projects and infrastructure
To be effective, efficient, collaborative, and ultimately successful in population suppression, future Burmese python research would require multi-year studies at the landscape scale, coordinated across multiple organizations with consistent effort-both in terms of funding and labor. One example is multi-year funding for collecting data to estimate key python demographic parameters (e.g., reproductive frequency, age-specific fecundity/survivorship) that are required to develop, evaluate, and ultimately maximize efficacy of control tools. Additionally, development of a facility in southern Florida for captive and small-scale manipulative trials could be useful to refine and optimize the application of control tools and explore techniques better equipped to handle low detection. Unlike the 5-ha enclosure built to enumerate a wild population of brown treesnakes and evaluate detection probability under various control approaches (e.g.,
Biologging tools to inform python behavior
Monitoring individual animals in situ has the potential to reveal important aspects of a species’ ecology such as seasonal and daily activity patterns, foraging strategies, or reproductive behaviors, and this approach may reveal vulnerabilities for control tool development or may improve python detection. The rise of electronic biologging devices in the last two decades has unlocked a wealth of information about animal space use, movements, and physiology (e.g.,
Technological advances in tags deployed in or on wildlife (i.e., biologging tags;
Although GPS-tracking of pythons has typically had low success (
Development of new control tools
With the rapid evolution of technology, new control tools could be applied to aid Burmese python management. Although research may continue to lead to better understanding, optimization, and implementation of existing control technologies to fully evaluate their impacts and transferability in new locations, investments in novel technologies have the potential to yield high rewards. One example is the rapidly growing field of genetic biocontrol, where genetic material is manipulated with the goal of decreasing the ability of an invasive species to thrive in the non-native environment. Genetic biocontrol methods have primarily been used to control disease-carrying pests in laboratory experiments (e.g., mosquitos,
An example of a potential gene drive target in Burmese pythons includes targeting and destroying the X chromosome (
Alternatively, non-replicating species-specific RNA interference (RNAi) technologies disrupt physiological functions in a target organism to reduce reproductive output or cause death through ingestion or topical application to a target organism (e.g.,
Although Burmese pythons are thought to primarily be ambush predators, they do engage in active foraging (see Feeding Strategy section), and if effective prey-scent attractants are developed, they may lure pythons into the area to feed on an RNAi bait and possibly into a trap. Development of control tools that concentrate individual pythons in time and space such as continued refinement of chemical attractants (see Pheromone section) or development of food-based attractants may be valuable tools for population suppression efforts. As with any novel biocontrol technology, this line of research is a longer-term investment strategy but has the potential to increase removal efficiency.
Technical aspects of the research
While potentially powerful, genetic biocontrol technologies are relatively novel in vertebrates (
Gene drive regulation and stakeholder engagement
Regulation on gene drive systems is a rapidly evolving topic governed at international, regional, and national levels (
Demography and genetic biocontrol
Developing and evaluating genetic biocontrol methods will require information on python life history parameters such as the prevalence of multiple paternity, sex or age specific survival rates, and variation in fecundity (see Demography section). Overall, there is little information on the size, population growth, or demographic structure of current python subpopulations. These data gaps may impede efforts to evaluate any applied genetic biocontrol tools and likewise prevent comprehensive understanding of how many biologically manipulated individuals would need to be released on the landscape to yield population declines. Recent and active progress is being made by USGS and others to fill these data gaps on life history knowledge (e.g.,
| Term | Definition |
|---|---|
| Active thermoregulation | Behaviors used by ectotherms such as basking, seeking shade, or altering body posture to change heating and cooling rates |
| Ambush predator | Sit-and-wait predators that capture or trap prey by stealth or luring behaviors |
| Biennial reproduction | Breeding every two years |
| Brumate | Metabolic adaptation in reptiles allowing them to conserve energy by becoming dormant during cold temperatures |
| Capture mark recapture | CMR; Capturing many organisms, marking them, releasing them back into the population, and then determining the probability of capture (i.e., ratio of marked to unmarked animals in the population) |
| CRISPR-Cas9 | Programmable protein ribonucleic acid complex used to target and edit specific DNA sequences |
| Critical thermal minimum | Low temperature at which mobility is lost; if temperatures continue to fall the lethal thermal minimum is reached, leading to death |
| Cryptic species | Visual, olfactory, or auditory concealment by an organism to avoid detection as a predation or antipredator strategy |
| Dietary generalist | Organism that consumes a wide variety of foods |
| Dispersal | Unidirectional movement of organisms away from place of birth |
| Early Detection & Distribution Mapping System | EDDMapS; Geospatial database of invasive species reports. https://www.eddmaps.org/ |
| Ectotherm | Organisms that rely on environmental heat sources to control body temperature |
| Energy budget | Quantification of the uptake of energy from the environment by an organism (feeding and digestion) how that energy is spent including for maintenance, development, growth, and reproduction |
| environmental DNA | eDNA; DNA released from an organism into the environment |
| Everglades virus | An alphavirus included in the Venezuelan equine encephalitis virus complex |
| Home range | Area used by an animal during normal activities such as foraging and mating, but excluding occasional exploratory excursions |
| Labyrinth morph/phenotype | Maze-like dorsal pattern selectively bred into the commercial snake/python trade |
| Lacey Act | United States law created in 1900 to restrict illegal wildlife trade, bar international importation of injurious species, and protect species at risk |
| Movement ecology | Subdiscipline of ecology connecting connects an animal’s movement path with environmental heterogeneity, available resources, navigational capacity, and its biology |
| Multiple paternity | More than one male siring a clutch or litter |
| Nidovirus | Diverse order of enveloped positive-strand RNA viruses that infect a range of vertebrate and invertebrate hosts and can cause serious diseases |
| Non-replicating, species-specific RNA interference | RNAi; Process where species-specific RNA molecules affect gene expression of key processes that impact an organism's fitness |
| Occupancy model | Approach to estimate probability that a species will occupy a site |
| Osmoregulation | Active regulation of an organism’s body fluids to maintain electrolyte concentrations (i.e., prevent fluids from becoming too dilute or concentrated) |
| Oviposit | Lay eggs |
| Parthenogenesis | Spontaneous development of an embryo from an unfertilized egg cell; reproducing without a male or stored sperm |
| Pentastomes | Parasitic arthropods requiring one or more intermediate hosts before completing its life cycle in a definitive host |
| Polymerase chain reaction | PCR; laboratory technique for rapidly producing (amplifying) millions to billions of copies of a specific segment of DNA for genetic analyses |
| Radiotelemetry | Attaching or implanting a transmitter to an animal and using a receiver and directional antenna to locate it over space and time |
| Scout snake | A radiotagged snake used to locate untagged snakes |
| Shivering thermogenesis | Generation of heat by repeated contraction of muscles |
| Serpentovirus | Also known as reptile nidovirus. See also: nidovirus. Virus that causes severe and often fatal respiratory disease, typically in captive snake species, especially pythons |
| Snake fungal disease | SFD; Infectious disease found in many snake species caused by the fungus Ophidiomyces ophidiicola |
| Spatial capture-recapture | SCR; Extension of capture-mark-recapture used to estimate population density from detections and subsequent redetections of individuals across space |
| Species or Individual detection | Chance that a species or individual will be detected during a survey, given that it is present at the location |
| Snout-vent length | SVL; Measurement of size taken from tip of nose to opening of cloaca, at base of tail |
| Survey | Ecological census conducted via a variety of methods to collect data on occupancy of habitats by an organism |
| Total length | TL; Measurement of size taken from tip of nose to tip of tail |
Conclusions
Burmese pythons in southern Florida represent one of the most intractable invasive-species management issues across the globe. The problem stems from a unique combination of inaccessible habitat with the cryptic and resilient nature of pythons that do very well in the subtropical environment of southern Florida, rendering them extremely difficult to detect. We have documented extensive direct alteration of the native food web as well as some aspects of the basic biology of these giant constrictors over the past two decades, while extensively exploring methods to capture and remove this damaging species (Table
Although a wide variety of techniques have been employed to catch pythons across southern Florida, many of these tools have not been evaluated rigorously, largely because of difficulty detecting pythons. Although rapid response to reports of individual pythons in new areas is ongoing, there have not been any concerted efforts aimed at suppression or eradication of python populations, even in limited areas. Cost-effective control methods and a better understanding of impacts on natural resources may help to inform application of limited resources and development of mitigation strategies. Because of individual heterogeneity in snake detection (e.g.,
Acknowledgements
We thank several entities for funding this manuscript, including the U.S Geological Survey (USGS) Greater Everglades Priority Ecosystem Science Program, USGS Biological Threats and Invasive Species Research Program, and the National Park Service. We thank Paul Andreadis for input during development of this manuscript, and Megan Arias, and Denise Gregoire-Lucente, and Adam Perez for assistance with manuscript preparation. We very much appreciate comments from Andrew Durso, Adam Perez, David Steen, Daniel Quinn, Eric Suarez, and two anonymous reviewers, which improved the manuscript. The findings and conclusions in this article are those of the author(s) and do not necessarily represent the views of the U.S. Fish and Wildlife Service. Any use of trade, firm, or product names is for descriptive purposes only and does not imply endorsement by the U.S. Government.
References
- Abatzoglou JT, Dobrowski SZ, Parks SA (2020) Multivariate climate departures have outpaced univariate changes across global lands. Scientific Reports 10(1): 1–9. https://doi.org/10.1038/s41598-020-60270-5
- Amburgey SM, Yackel Adams AA, Gardner B, Lardner B, Knox AJ, Converse SJ (2021) Tools for increasing visual encounter probabilities for invasive species removal: A case study of brown treesnakes. NeoBiota 70: 107–122. https://doi.org/10.3897/neobiota.70.71379
- Anderson DR (2001) The need to get the basics right in wildlife field studies. Wildlife Society Bulletin 2(29): 1294–1297.
- Anderson GE, Ridgley FN, Josimovich JM, Reed RN, Falk BG, Yackel Adams AA, Currylow AF (2022) Egg retention in wild-caught Python bivittatus in the Greater Everglades Ecosystem, Florida, USA. The Herpetological Journal 32(3): 109–113. https://doi.org/10.33256/32.3.109113
- Andreadis PT (2011) Python molurus bivittatus (Burmese Python). Reproducing Population. Herpetological Review 42: 302–303.
- Andreadis PT, Bartoszek IA, Prokop-Ervin C, Pittman SE (2018) Drymarchon kolpobasileus (Gulf Coast indigo snake) and Python bivittatus (Burmese python) predator/prey interaction. Herpetological Review 49: 341–342.
- Andrew AL, Perry BW, Card DC, Schield DR, Ruggiero RP, McGaugh SE, Choudhary A, Secor SM, Castoe TA (2017) Growth and stress response mechanisms underlying post-feeding regenerative organ growth in the Burmese python. BMC Genomics 18(1): 18. https://doi.org/10.1186/s12864-017-3743-1
- ATSDR (1999) Agency for Toxic Substances and Disease Registry: Mercury-ToxFAQs. https://www.atsdr.cdc.gov/toxfaqs/tfacts46.pdf
- Avery ML, Engeman RM, Keacher KL, Humphrey JS, Bruce WE, Mathies TC, Mauldin RE (2010) Cold weather and the potential range of invasive Burmese pythons. Biological Invasions 12(11): 3649–3652. https://doi.org/10.1007/s10530-010-9761-4
- Avery ML, Humphrey JS, Keacher KL, Bruce EW (2014) Detection and removal of invasive Burmese pythons: Methods development update. Preceedings of the Vertebrate Pest Conference 26: 145–148. https://doi.org/10.5070/V426110362
- Bailey LL, MacKenzie DI, Nichols JD (2014) Advances and applications of occupancy models. Methods in Ecology and Evolution 5(12): 1269–1279. https://doi.org/10.1111/2041-210X.12100
- Bajer PG, Chizinski CJ, Sorensen PW (2011) Using the Judas technique to locate and remove wintertime aggregations of invasive common carp. Fisheries Management and Ecology 18(6): 497–505. https://doi.org/10.1111/j.1365-2400.2011.00805.x
- Baker RH, Newman CC, Wilke F (1945) Food habits of the Raccoon in Eastern Texas. The Journal of Wildlife Management 9(1): 45–48. https://doi.org/10.2307/3795945
- Barker DG, Barker TM (2008) The distribution of the Burmese python, Python molurus bivittatus. Bulletin of the Chicago Herpetological Society 43: 33–38.
- Bartoszek IA (2022) Record-breaking python caught in Florida measured in at 17.7 feet, 215 pounds: ABC News. [Accessed 9/21/2022] https://abc7news.com/python-longest-nat-geo-florida/11984390/
- Bartoszek IA, Andreadis PT, Prokop-Ervin C, Curry G, Reed RN (2018a) Python bivattatus (Burmese python) and Gopherus polyphemus (gopher tortoise). Habitat use, breeding aggregation, and interspecific interaction. Herpetological Review 49: 353–3584.
- Bartoszek IA, Andreadis PT, Prokopervin C, Patel M, Reed RN (2018b) Python bivittatus (Burmese python). Diet and Prey Size. Herpetological Review 49: 139–140.
- Bartoszek IA, Hendricks MB, Easterling IC, Andreadis PT (2018c) Python bivittatus (Burmese Python). Dispersal/Marine Incursion. Herpetological Review 49: 554–555.
- Bartoszek IA, Easterling IC, King K, Andreadis PT (2020) Python bivittatus (Burmese python). Aberrant pattern. Herpetological Review 51: 155–156.
- Bartoszek IA, Anderson GE, Easterling IC, Josimovich JM, Furst A, Ridgley FN, Fitzgerald AL, Yakel Adams AA, Currylow AF (2021a) Agkistrodon conanti and Python bivittatus. Diet and Predation. Herpetological Review 52: 860–862.
- Bartoszek IA, Smith BJ, Reed RN, Hart KM (2021b) Spatial ecology of invasive Burmese pythons in southwestern Florida. Ecosphere 12(6): 12. https://doi.org/10.1002/ecs2.3564
- Bauder JM, Allen ML, Benson TJ, Miller CA, Stodola KW (2021) An approach for using multiple indices for monitoring long-term trends of mesopredators at broad spatial scales. Biodiversity and Conservation 30(12): 3529–3547. https://doi.org/10.1007/s10531-021-02259-8
- Bauer AM, Günther R (2013) Origin and identity of the von Borcke collection of amphibians and reptiles in the Museum für Naturkunde in Berlin: A cache of Seba specimens? Zoosystematics and Evolution 89(1): 167–185. https://doi.org/10.1002/zoos.201300005
- Beaupré SJ, Montgomery CE (2007) The meaning and consequences of foraging mode in snakes. In: Reilly SM, et al. (Eds) Lizard Ecology. Cambridge University Press, New York, 334–367. https://doi.org/10.1017/CBO9780511752438.013
- Beebe SC, Howell TJ, Bennett PC (2016) Using scent detection dogs in conservation settings: A review of scientific literature regarding their selection. Frontiers in Veterinary Science 3: 1–13. https://doi.org/10.3389/fvets.2016.00096
- Beng KC, Corlett RT (2020) Applications of environmental DNA (eDNA) in ecology and conservation: Opportunities, challenges and prospects. Biodiversity and Conservation 29(7): 2089–2121. https://doi.org/10.1007/s10531-020-01980-0
- Bhupathy S (1995) Distribution of Python molurus bivittatus in India. Cobra 21: 2–5.
- Bhupathy S, Vijayan VS (1989) Status, distribution and general ecology of the Indian python Python molurus molurus linn. in Keoladeo National Park, Bharatpur, Rajasthan. Journal of the Bombay Natural History Society 86: 381–387.
- Blair WF (1936) The Florida Marsh Rabbit. Journal of Mammalogy 17(3): 197–207. https://doi.org/10.2307/1374414
- Blair WF (1940) Notes on home ranges and populations of the short-tailed shrew. Ecology 21(2): 284–288. https://doi.org/10.2307/1930504
- Blundell AG, Mascia MB (2005) Discrepancies in reported levels of international wildlife trade. Conservation Biology 19(6): 2020–2025. https://doi.org/10.1111/j.1523-1739.2005.00253.x
- Boback SM, Snow RW, Hsu T, Peurach SC, Dove CJ, Reed RN (2016) Supersize me: remains of three white-tailed deer (Odocoileus virginianus) in an invasive Burmese python (Python molurus bivittatus) in Florida. https://doi.org/10.3391/bir.2016.5.4.02
- Bond WJ (1994) Keystone species. In: Schulze ED, Mooney HA (Eds) Biodiversity and ecosystem Function. Springer Verlag, Germany, 237–253. https://doi.org/10.1007/978-3-642-58001-7_11
- Bonneau M, Johnson FA, Romagosa CM (2016) Spatially explicit control of invasive species using a reaction-diffusion model. Ecological Modelling 337: 15–24. https://doi.org/10.1016/j.ecolmodel.2016.05.013
- Bonnet X, Naulleau G, Shine R, Lourdais O (2000) Reproductive versus ecological advantages to larger body size in female snakes, Vipera aspis. Oikos 89(3): 509–518. https://doi.org/10.1034/j.1600-0706.2000.890310.x
- Booth W, Schuett GW, Ridgway A, Buxton DW, Castoe TA, Bastone G, Bennett C, Mcmahan W (2014) New insights on facultative parthenogenesis in pythons. Biological Journal of the Linnean Society. Linnean Society of London 112(3): 461–468. https://doi.org/10.1111/bij.12286
- Bowler JK (1977) Longevity of reptiles and amphibians in North American collections. Society for the Study of Amphibians and Reptiles and the Philidelphia Herpetological Society.
- Boyce MS, Sinclair ARE, White GC (1999) Seasonal Compensation of Predation and Harvesting 87: 419–426. https://doi.org/10.2307/3546808
- Branch B, Erasmus H (1984) Captive breeding of pythons in south africa, including details of an interspecific hybrid (Python sebae natalensis × Python molurus bivittatus). Journal of the Herpetological Association of Africa 30(1): 1–10. https://doi.org/10.1080/04416651.1984.9650132
- Brashears JA, DeNardo DF (2013) Revisiting python Thermogenesis: Brooding Burmese pythons (Python bivittatus) cue on body, not clutch, temperature. Journal of Herpetology 47(3): 440–444. https://doi.org/10.1670/12-050
- Bravington MV, Skaug HJ, Anderson EC (2016) Close-kin mark-recapture. Statistical Science 31(2): 259–274. https://doi.org/10.1214/16-STS552
- Brooks JE, Savarie PJ, Johnston JJ (1998) The oral and dermal toxicity of selected chemicals to brown tree snakes (Boiga irregularis). Wildlife Research 25(4): 427–435. https://doi.org/10.1071/WR97035
- Bruton MJ (2013) Arboreality, excavation, and active foraging: Novel observations of radiotracked woma pythons Aspidites ramsayi. Memoirs of the Queensland Museum 56: 313–329.
- Buckley YM, Hinz HL, Matthies D, Rees M (2001) Interactions between density-dependent processes, population dynamics and control of an invasive plant species, Tripleurospermum perforatum (scentless chamomile). Ecology Letters 4(6): 551–558. https://doi.org/10.1046/j.1461-0248.2001.00264.x
- Bullock JM, Kenward RE, Hails RS [Eds] (2002) Dispersal Ecology. Blackwell Science, Oxford.
- Burgdorf SJ, Rudolph DC, Conner RN, Saenz D, Schaefer RR (2005) A successful trap design for capturing large terrestrial snakes. Herpetological Review 36: 421–424.
- Burkett-Cadena ND, Blosser EM, Loggins AA, Valente MC, Long MT, Campbell LP, Reeves LE, Bargielowski I, McCleery RA (2021) Invasive Burmese pythons alter host use and virus infection in the vector of a zoonotic virus. Communications Biology 4(1): 1–11. https://doi.org/10.1038/s42003-021-02347-z
- Burnham KP, Anderson DR (1984) Tests of compensatory vs. Additive hypotheses of mortality in Mallards. Ecology 65(1): 105–112. https://doi.org/10.2307/1939463
- Burt WH (1943) Territoriality and home range concepts as applied to mammals. Page Journal of Mammalogy 24(3): 346–352. https://doi.org/10.2307/1374834
- Butterfield BP, Meshaka Jr WE, Guyer C (1997) Nonindigenous amphibians and reptiles. Strangers in Paradise. Impact and Management of Nonindiginous Species in Florida, 123–138.
- Cagnacci F, Boitani L, Powell RA, Boyce MS (2010) Animal ecology meets GPS-based radiotelemetry: A perfect storm of opportunities and challenges. Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences 365(1550): 2157–2162. https://doi.org/10.1098/rstb.2010.0107
- Cam E (2009) Modeling demographic processes in marked populations. In: Thomson DL, et al. (Eds) Environmental and Ecological Statistics. Volume 3. Springer, 83–129. https://doi.org/10.1007/978-0-387-78151-8_5
- Campbell KJ, Baxter GS, Murray PJ, Coblentz BE, Donlan CJ, Carrion VG (2005) Increasing the efficacy of Judas goats by sterilisation and pregnancy termination. Wildlife Research 32(8): 737–743. https://doi.org/10.1071/WR05033
- Campbell EW, Yackel Adams AA, Converse SJ, Fritts TH, Rodda GH (2012) Do predators control prey species abundance? An experimental test with brown treesnakes on Guam. Ecology 93(5): 1194–1203. https://doi.org/10.1890/11-1359.1
- Campbell JA, Smith EN, Hall AS (2018) Caudals and Calyces: The Curious Case of a Consumed Chiapan Colubroid. Journal of Herpetology 52(4): 459–472. https://doi.org/10.1670/18-042
- Card DC, Perry BW, Adams RH, Schield DR, Young AS, Andrew AL, Jezkova T, Pasquesi GIM, Hales NR, Walsh MR, Rochford MR, Mazzotti FJ, Hart KM, Hunter ME, Castoe TA (2018) Novel ecological and climatic conditions drive rapid adaptation in invasive Florida Burmese pythons. Molecular Ecology 27(23): 4744–4757. https://doi.org/10.1111/mec.14885
- Castoe TA, Jason de Koning A, Hall KT, Yokoyama KD, Gu W, Smith EN, Uetz P, Ray DA, Dobry J, Bogden R, Mackessy SP, Bronikowski AM, Secor SM, Pollock DD (2011) The importance of snakes, and the Burmese python, as model organisms Sequencing the genome of the Burmese python (Python molurus bivittatus) as a model for studying extreme adaptations in snakes. Page Genome Biology 12: 406. https://doi.org/10.1186/gb-2011-12-7-406
- Castoe TA, De Koning APJ, Hall KT, Card DC, Schield DR, Fujita MK, Ruggiero RP, Degner JF, Daza JM, Gu W, Reyes-Velasco J, Shaney KJ, Castoe JM, Fox SE, Poole AW, Polanco D, Dobry J, Vandewege MW, Li Q, Schott RK, Kapusta A, Minx P, Feschotte C, Uetz P, Ray DA, Hoffmann FG, Bogden R, Smith EN, Chang BSW, Vonk FJ, Casewell NR, Henkel CV, Richardson MK, Mackessy SP, Bronikowsi AM, Yandell M, Warren WC, Secor SM, Pollock DD (2013) The Burmese python genome reveals the molecular basis for extreme adaptation in snakes. Proceedings of the National Academy of Sciences of the United States of America 110(51): 20645–20650. https://doi.org/10.1073/pnas.1314475110
- Caswell H (2001) Matrix population models: Construction, analysis, and interpretation. 2nd Edn. Sinauer Associates, Inc., Sunderland, MA.
- Caudill G, Onorato D, Cunningham MW, Caudill D, Leone EH, Smith L, Jansen D (2019) Temporal trends in Florida panther food habits. Human-Wildlife Interactions 13: 87–97.
- Center TD, Purcell MF, Pratt PD, Rayamajhi MB, Tipping PW, Wright SA, Dray Jr FA (2012) Biological control of Melaleuca quinquenervia: An Everglades invader. BioControl 57(2): 151–165. https://doi.org/10.1007/s10526-011-9390-6
- Christoffersen ML, De Assis JE (2013) A systematic monograph of the Recent Pentastomida, with a compilation of their hosts. Zoölogische Mededeelingen 87: 206.
- Christy MT, Yackel Adams AA, Rodda GH, Savidge JA, Tyrrell CL (2010) Modelling detection probabilities to evaluate management and control tools for an invasive species. Journal of Applied Ecology 47(1): 106–113. https://doi.org/10.1111/j.1365-2664.2009.01753.x
- CITES (2020) CITES trade statistics derived from the CITES Trade Database. UNEP World Conservation Monitoring Centre, Cambridge, UK. https://trade.cites.org/en/cites_trade/#
- Clark B (1996) Python Color and Pattern Morphs. Reptiles, 36–62.
- Clark L, Savarie PJ, Shivik JA, Breck SW, Dorr BS (2012) Efficacy, effort, and cost comparisons of trapping and acetaminophen-baiting for control of brown treesnakes on Guam. Human-Wildlife Interactions 6: 222–236.
- Clarke AL, Dalrymple GH (2003) $ 7.8 Billion for Everglades Restoration: Why Do Environmentalists Look so Worried? Population and Environment 24(6): 541–569. https://doi.org/10.1023/A:1025030832054
- Claunch NM, Nix E, Royal AE, Burgos LP, Corn M, DuBois PM, Ivey KN, King EC, Rucker KA, Shea TK, Stepanek J, Vansdadia S, Taylor EN (2021) Body size impacts critical thermal maximum measurements in lizards. Journal of Experimental Zoology. Part A, Ecological and Integrative Physiology 335(1): 96–107. https://doi.org/10.1002/jez.2410
- Claunch NM, Bartoszek IA, Tillis S, Stacy NI, Ossiboff RJ, Oakey S, Schoenle LA, Wellehan JFX, Romagosa CM (2022) Physiological effects of capture and short-term captivity in an invasive snake species, the Burmese python (Python bivittatus) in Florida. Comparative Biochemistry and Physiology. Part A, Molecular & Integrative Physiology 267: 111162. https://doi.org/10.1016/j.cbpa.2022.111162
- Clercq PD (1988) Breeding results: Python molurus bivittatus. Litteratura Serpentium 8(1): 50.
- Coffey LL, Crawford C, Dee J, Miller R, Freier J, Weaver SC (2006) Serologic Evidence of Widespread Everglades Virus Activity in Dogs, Florida. Emerging Infectious Diseases 12(12): 1873–1879. https://doi.org/10.3201/eid1212.060446
- Collins T, Freeman B, Snow S (2008) Genetic characterization of populations of the nonindigenous Burmese python in Everglades National Park. Page final report for the South Florida water management district prepared by the department of biological sciences, Florida International University, Miami, FL.
- Conant R (1938) The Reptiles of Ohio. Page The American Midland Naturalist. The University of Notre Dame 20(1): 1–200. https://doi.org/10.2307/2485190
- Congdon JD, Dunham AE, Tinkle DW (1982) Energy budgets and life histories of reptiles. In: Gans C, Pough FH (Eds) Biology of the Reptilia. Academic Press, London, 233–272.
- Congdon JD, Greene JL, Gibbons JW (1986) Biomass of Freshwater Turtles: A Geographic Comparison. American Midland Naturalist 115(1): 165–173. https://doi.org/10.2307/2425846
- Corn JL, Mertins JW, Hanson B, Snow S (2011) First reports of ectoparasites collected from wild-caught exotic reptiles in Florida. Journal of Medical Entomology 48(1): 94–100. https://doi.org/10.1603/ME10065
- Cowles RB, Bogert CM (1944) A preliminary study of the thermal requirements of desert reptiles. Bulletin of the American Museum of Natural History 83: 261–296.
- Cox CL, Secor SM (2007) Effects of meal size, clutch, and metabolism on the energy efficiencies of juvenile Burmese pythons, Python molurus. Comparative biochemistry and physiology. Part A, Molecular & Integrative Physiology 148(4): 861–868. https://doi.org/10.1016/j.cbpa.2007.08.029
- Cullen JA, Poli CL, Fletcher Jr RJ, Valle D (2022) Identifying latent behavioural states in animal movement with M4, a nonparametric Bayesian method. Methods in Ecology and Evolution 13(2): 432–446. https://doi.org/10.1111/2041-210X.13745
- Cupp AR, Brey MK, Calfee RD, Chapman DC, Erickson R, Fischer J, Fritts AK, George AE, Jackson PR, Knights BC, Saari GN, Kočovský PM (2021) Emerging control strategies for integrated pest management of invasive carps. Journal of Vertebrate Biology 70(4): 21057. https://doi.org/10.25225/jvb.21057
- Currylow AF, Falk BG, Yackel Adams AA, Romagosa C, Josimovich JM, Rochford M, Cherkiss M, Nafus MG, Hart K, Mazzotti FM, Snow R, Reed RN (2022b) Size distribution and reproductive phenology of the invasive Burmese python (Python molurus bivittatus) in the Greater Everglades Ecosystem, Florida, USA. NeoBiota 78: 129–158. https://doi.org/10.3897/neobiota.78.93788
- Currylow AF, Fitzgerald AL, Goetz MT, Draxler JL, Anderson GE, McCollister MF, Romagosa CM, Yackel Adams AA (In press) Natives bite back: Depredation and mortality of invasive juvenile Burmese pythons (Python bivittatus) in the Greater Everglades Ecosystem. Management of Biological Invasions.
- Currylow AF, McCollister MF, Anderson GE, Josimovich JM, Fitzgerald AL, Romagosa CM, Yackel Adams AA (2022a) Face‐off: Novel depredation and nest defense behaviors between an invasive and a native predator in the Greater Everglades Ecosystem, Florida, USA. Ecology and Evolution 12(2): 1–6. https://doi.org/10.1002/ece3.8639
- Currylow AF, Falk BG, Yackel Adams AA, Romagosa CM, Josimovich JM, Rochford MR, Cherkiss MS, Nafus MG, Hart KM, Mazzotti FJ, Snow RW, Reed RN (2022c) Size distribution and reproductive data of the invasive Burmese python (Python molurus bivittatus) in the Greater Everglades Ecosystem, Florida, USA, 1995–2021: U.S. Geological Survey data release. NeoBiota 78: 129–158. https://doi.org/10.3897/neobiota.78.93788
- D’Acunto LE, Pearlstine L, Romañach SS (2021) Joint species distribution models of Everglades wading birds to inform restoration planning. PLoS ONE 16(1): e0245973. https://doi.org/10.1371/journal.pone.0245973
- Daniel JC (2002) The book of Indian reptiles and amphibians. Bombay Natural History Society.
- Davies PMC, Bennett EL (1981) Non-acclimatory latitude-dependent metabolic adaptation to temperature in juvenile natricine snakes. Journal of Comparative Physiology. B, Biochemical, Systemic, and Environmental Physiology 142(4): 489–494. https://doi.org/10.1007/BF00688980
- Davis S, Ogden JC (1994) Everglades: the ecosystem and its restoration. CRC Press, 860 pp.
- Davis AJ, Hooten MB, Miller RS, Farnsworth ML, Lewis J, Moxcey M, Pepin KM (2016) Inferring invasive species abundance using removal data from management actions. Ecological Applications 26(7): 2339–2346. https://doi.org/10.1002/eap.1383
- Davis AJ, Farrar R, Jump B, Hall P, Guerrant T, Pepin KM (2021) An efficient method of evaluating multiple concurrent management actions on invasive populations. bioRxiv, 1–22. https://doi.org/10.1101/2021.07.28.452079
- de Queiroz KD (1999) The General Lineage Concept of Species and the Defining Properties of the Species Category. In: Wilson RA (Ed.) Species: New Interdisciplinary Essays. MIT, 49–89.
- Delibes-Mateos M, Delibes M, Ferreras P, Villafuerte R (2008) Key role of European rabbits in the conservation of the western Mediterranean Basin hotspot. Conservation Biology 22(5): 1106–1117. https://doi.org/10.1111/j.1523-1739.2008.00993.x
- Diffie S, Miller J, Murray K (2010) Laboratory observations of red imported fire ant (Hymenoptera: Formicidae) predation on reptilian and avian eggs. Journal of Herpetology 44(2): 294–296. https://doi.org/10.1670/08-282.1
- Dodd CK [Ed.] (2016) Reptile ecology and conservation. A handbook of techniques. Oxford University Press, New York, NY, 512 pp. https://doi.org/10.1093/acprof:oso/9780198726135.001.0001
- Doherty TS, Glen AS, Nimmo DG, Ritchie EG, Dickman CR (2016) Invasive predators and global biodiversity loss. Proceedings of the National Academy of Sciences of the United States of America 113(40): 11261–11265. https://doi.org/10.1073/pnas.1602480113
- Dorazio RM, Erickson RA (2018) EDNAOCCUPANCY: An R package for multiscale occupancy modelling of environmental DNA data. Molecular Ecology Resources 18(2): 368–380. https://doi.org/10.1111/1755-0998.12735
- Dorazio RM, Jelks HL, Jordan F (2005) Improving removal-based estimates of abundance by sampling a population of spatially distinct subpopulations. Biometrics 61(4): 1093–1101. https://doi.org/10.1111/j.1541-0420.2005.00360.x
- Dorazio RM, Mukherjee B, Zhang L, Ghosh M, Jelks HL, Jordan F (2008) Modeling unobserved sources of heterogeneity in animal abundance using a Dirichlet process prior. Biometrics 64(2): 635–644. https://doi.org/10.1111/j.1541-0420.2007.00873.x
- Dorcas ME, Willson JD (2009) Innovative methods for studies of snake ecology and conservation. Snakes: ecology and conservation. 1st edn. Cornell University Press, Comstock Publishing Associates, 5–37. https://doi.org/10.7591/9780801459092-005
- Dorcas ME, Willson JD (2013) Hidden Giants: Problems Associated with Studying Secretive Invasive Pythons. In: Lutterschmidt WI (Ed.) Reptiles in research: Investigations of ecology, physiology, and behavior from desert to sea. Nova Biomedical, 367–385.
- Dorcas ME, Willson JD, Gibbons JW (2011) Can invasive Burmese pythons inhabit temperate regions of the southeastern United States? Biological Invasions 13(4): 793–802. https://doi.org/10.1007/s10530-010-9869-6
- Dorcas ME, Willson JD, Reed RN, Snow RW, Rochford MR, Miller MA, Meshaka Jr WE, Andreadis PT, Mazzotti FJ, Romagosa CM, Hart KM (2012) Severe mammal declines coincide with proliferation of invasive Burmese pythons in Everglades National Park. Proceedings of the National Academy of Sciences of the United States of America 109(7): 2418–2422. https://doi.org/10.1073/pnas.1115226109
- Doughty P (1994) Critical thermal minima of garter snakes (Thamnophis) depend on species and body size. Copeia 1994(2): 537–540. https://doi.org/10.2307/1447008
- Dove CJ, Snow RW, Rochford MR, Mazzotti FJ (2011) Birds consumed by the invasive Burmese python (Python molurus bivittatus) in Everglades National Park, Florida, USA. The Wilson Journal of Ornithology 123(1): 126–131. https://doi.org/10.1676/10-092.1
- Dove CJ, Reed RN, Snow RW (2012) Consumption of bird eggs by invasive Burmese Pythons in Florida. Reptiles & Amphibians: Conservation and Natural History 19(1): 64–66. https://doi.org/10.17161/randa.v19i1.13848
- Driggers R, Furxhi O, Vaca G, Reumers V, Vazimali M, Short R, Agrawal P, Lambrechts A, Charle W, Vunckx K, Arvidson C (2019) Burmese python target reflectivity compared to natural Florida foliage background reflectivity. Applied Optics 58(13): D98–D104. https://doi.org/10.1364/AO.58.000D98
- Dunson WA, Mazzotti FJ (1989) Salinity as a limiting factor in the distribution of reptiles in Florida Bay: A theory for the estuarine origin of marine snakes and turtles. Bulletin of Marine Science 44: 229–244.
- Durso AM, Willson JD, Winne CT (2011) Needles in haystacks: Estimating detection probability and occupancy of rare and cryptic snakes. Biological Conservation 144(5): 1508–1515. https://doi.org/10.1016/j.biocon.2011.01.020
- Easterling IC, Bartoszek IA (2019) Python bivittatus (Burmese python). Maximum male size. Herpetological Review 50: 399–400.
- EDDMapS (2018) Early Detection and Distribution Mapping System (EDDMAPS). The University of Georgia: Center for Invasive Species and Ecosystem Health. [Accessed September 2020]
- Enge KM (1997) Use of Silt Fencing and Funnel Traps for Drift Fences. Herpetological Review 28: 30–31.
- Enge KM, Berish JD, Moler P, Thomas TM (2012) What the world needs is a better gopher tortoise trap. Herpetological Review 43: 574–578.
- Engeman RM (2005) Indexing principles and a widely applicable paradigm for indexing animal populations. Wildlife Research 32(3): 203–210. https://doi.org/10.1071/WR03120
- Engeman RM, Kennedy M, Constantin BU, Christie ML, Hall PT (2009) Ctenosaura similis (Black Spiny-Tailed Iguana), Coluber constrictor priapus (Southern Black Racer). NON-PREDATORY KILLING. Herpetological Review 40: 84–85.
- Engeman R, Avery ML, Jacobson E (2014) Weighing empirical and hypothetical evidence for assessing potential invasive species range limits: A review of the case of Burmese pythons in the USA. Environmental Science and Pollution Research International 21(20): 11973–11978. https://doi.org/10.1007/s11356-014-3173-4
- Ernst CH, Lovich JE (1999) Turtles of the United States and Canada. Johns Hopkins University Press, Baltimore, MD, USA.
- Eversole CB, Henke SE, Ballard BM, Powell RL (2014) Duration of marking tags on American Alligators (Alligator mississippiensis). Herpetological Review 45: 223–226.
- Faber NR, McFarlane GR, Gaynor RC, Pocrnic I, Whitelaw CBA, Gorjanc G (2021) Novel combination of CRISPR-based gene drives eliminates resistance and localises spread. Scientific Reports 11(1): 3719. https://doi.org/10.1038/s41598-021-83239-4
- Falk BG, Reed RN (2015) Challenges to a molecular approach to prey identification in the Burmese python, Python molurus bivittatus. PeerJ 3: e1445. https://doi.org/10.7717/peerj.1445
- Falk BG, Snow RW, Reed RN (2016) Prospects and limitations of citizen science in invasive species management: A case study with Burmese pythons in Everglades National Park. Southeastern Naturalist (Steuben, ME) 15(sp8): 89–102. https://doi.org/10.1656/058.015.sp806
- Falk BG, Snow RW, Reed RN (2017) A validation of 11 body-condition indices in a giant snake species that exhibits positive allometry. PLoS ONE 12(7): 1–20. https://doi.org/10.1371/journal.pone.0180791
- Fantle-Lepczyk JE, Haubrock PJ, Kramer AM, Cuthbert RN, Turbelin AJ, Crystal-Ornelas R, Diagne C, Courchamp F (2022) Economic costs of biological invasions in the United States. The Science of the Total Environment 806: 151318. https://doi.org/10.1016/j.scitotenv.2021.151318
- Farrell TM, Agugliaro J, Walden HDS, Wellehan JX, Childress AL, Lind CM (2019) Spillover of Pentastoma Parasites from Invasive Burmese Pythons (Python bivittatus) to Pygmy Rattlesnakes (Sistrurus miliarius), Extending Parasite Range in Florida, USA. Herpetological Review 50: 73–76.
- Ficetola GF, Miaud C, Pompanon F, Taberlet P (2008) Species detection using environmental DNA from water samples. Biology Letters 4(4): 423–425. https://doi.org/10.1098/rsbl.2008.0118
- Finkl CW, Makowski C (2017) The Florida everglades: An overview of alteration and restoration. In: Finkl CW, Makowski C (Eds) Coastal Wetlands: Alteration and Remediation. Springer, 512 pp. https://doi.org/10.1007/978-3-319-56179-0
- Fish and Wildlife Service (2015) Injurious Wildlife Species; Listing Three Anaconda Species and One Python Species as Injurious Reptiles. Federal Register, 50 CFR Part 16, 12702–12745.
- Fitch HS (1951) A simplified type of funnel trap for reptiles. Herpetologica 7: 77–80.
- Fitch HS (1963) Natural History of the Black Rat Snake (Elaphe o. obsoleta) in Kansas. Copeia 1963(4): 649–658. https://doi.org/10.2307/1440967
- Fitzgerald LA (2012) Finding and capturing reptiles. Reptile Biodiversity: Standard Methods for Inventory and Monitoring, 77–88.
- Fitzgerald AL, Josimovich JM, Robinson CJ, Reed RN, Currylow AF (2021) Identifying negative sentiment polarity in the Judas technique. Conservation Science and Practice 3(11): 1–7. https://doi.org/10.1111/csp2.532
- Frederick P, Gawlik DE, Ogden JC, Cook MI, Lusk M (2009) The White Ibis and Wood Stork as indicators for restoration of the everglades ecosystem. Ecological Indicators 9(6): S83–S95. https://doi.org/10.1016/j.ecolind.2008.10.012
- Fritts TH, Rodda GH (1998) The role of introduced species in the degradation of island ecosystems: A case history of Guam. Annual Review of Ecology and Systematics 29(1): 113–140. https://doi.org/10.1146/annurev.ecolsys.29.1.113
- Frýdlová P, Mrzílková J, Šeremeta M, Kremen J, Dudák J, Zemlicka J, Nemec P, Velenský P, Moravec J, Koleška D, Zahradníckova V, Jirásek T, Kodym P, Frynta D, Zach P (2019) Universality of indeterminate growth in lizards rejected: The micro-CT reveals contrasting timing of growth cartilage persistence in iguanas, agamas, and chameleons. Scientific Reports 9(1): 1–14. https://doi.org/10.1038/s41598-019-54573-5
- Frye FL, Mader DR (1985) Precocious sexual maturity in a juvenile male python. The Journal of Wildlife Management 16: 69–72. https://doi.org/10.2307/20094747
- FWC (2017) Final Report Submitted to the Florida Fish and Wildlife Conservation Commission TA-A3017: Assessment of Python Detector Dogs and eDNA June 2017.
- FWC (2022) Florida Fish and Wildlife Conservation Commission. PATRIC: Python Action Team Removing Invasive Constrictors. https://myfwc.com/wildlifehabitats/nonnatives/python/action-team/
- Galizi R, Hammond A, Kyrou K, Taxiarchi C, Bernardini F, O’Loughlin SM, Papathanos P-A, Nolan T, Windbichler N, Crisanti A (2016) A CRISPR-Cas9 sex-ratio distortion system for genetic control. Scientific Reports 6(1): 31139. https://doi.org/10.1038/srep31139
- Gamble T, Castoe TA, Nielsen SV, Banks JL, Card DC, Schield DR, Schuett GW, Booth W (2017) The Discovery of XY Sex Chromosomes in a Boa and Python. Current Biology 27(14): 2148–2153. https://doi.org/10.1016/j.cub.2017.06.010
- Gati EV, Dalaba JR, Miller MA, Humphrey JS, Kluever BM, Gibble R, Mazzotti FJ (2020) Assessment of Python Trapping within the Everglades Region Using a Patented Large Reptile Trap. Final Report to the Florida Fish and Wildlife Conservation Commission. Contract Number 13416-A3045.
- Gawlik DE (2002) The effects of prey availability on the numerical response of wading birds. Ecological Monographs 72(3): 329–346. https://doi.org/10.1890/0012-9615(2002)072[0329:TEOPAO]2.0.CO;2
- Gillingham JC, Chambers JA (1982) Courtship and pelvic spur use in the Burmese python, Python molurus bivittatus. Copeia 1982(1): 193–196. https://doi.org/10.2307/1444292
- Gilmour CC, Riedel GS, Ederington MC, Bell JT, Benoit JM (1998) Methylmercury concentrations and Production Rates across a. Trophic Gradient in the Northern Everglades. Biogeochemistry 40(2/3): 327–345. https://doi.org/10.1023/A:1005972708616
- Glorioso B, Bartoszek IA, Lorch JM (2020) Low-level detection of SFD-causing Ophidiomyces on Burmese pythons in southwest Florida, with confirmation of the pathogen on co-occurring native snakes. Herpetological Review 51: 245–247.
- Godfrey ST, Squires MA, Metzger III EF, Mazzotti FJ, Darling R, Muhly R (2021) Crocodylus acutus (American crocodile). Interspecific Interaction. Herpetological Review 52: 641–642.
- Goel VK, Goel B, Durso AM (2017) Python molurus. Predation. Natural History Notes 48: 866–867.
- Goris RC, Nakano M, Atobe Y, Funakoshi K (2007) The infrared sight of boas and pythons. In: Henderson RW, Powell R (Eds) Biology of the Boas and Pythons. Eagle Mountain Publishing.
- Govindarajulu P, Altwegg R, Anholt BR (2005) Matrix model investigation of invasive species control: Bullfrogs on Vancouver island. Ecological Applications 15(6): 2161–2170. https://doi.org/10.1890/05-0486
- Grace MS, Matushita A (2007) Neural correlates of complex behavior: vision and infrared imaging in boas and pythons. Biology of the boas and pythons. Eagle Mountain Publishing, 271–285.
- Gragg JE, Rodda H, Gordon A (2007) Response of brown treesnakes to reduction of their rodent prey. The Journal of Wildlife Management 71(7): 2311–2317. https://doi.org/10.2193/2006-444
- Greene DU, Potts JM, Duquesnel JG, Snow RW (2007) Python molurus bivittatus (Burmese python). Herpetological Review 38: 355.
- Groombridge B, Luxmoore R (1991) Pythons in South-east Asia: a review of distribution, status, and trade in three selected species. CITES, Lausanne, Switzerland.
- Groot TVM, Bruins E, Breeuwer JAJ (2003) Molecular genetic evidence for parthenogenesis in the Burmese python, Python molurus bivittatus. Heredity 90(2): 130–135. https://doi.org/10.1038/sj.hdy.6800210
- Guillera-Arroita G, Lahoz-Monfort JJ, van Rooyen AR, Weeks AR, Tingley R (2017) Dealing with false-positive and false-negative errors about species occurrence at multiple levels. Methods in Ecology and Evolution 8(9): 1081–1091. https://doi.org/10.1111/2041-210X.12743
- Guthrie JE (1932) Snakes versus birds; birds versus snakes. The Wilson Bulletin 44: 88–113.
- Hahn MA, Rieseberg LH (2017) Genetic admixture and heterosis may enhance the invasiveness of common ragweed. Evolutionary Applications 10(3): 241–250. https://doi.org/10.1111/eva.12445
- Hallac D, Kline J, Sadle J, Bass S, Ziegler T, Snow S (2010) Preliminary effects of the January 2010 cold weather on flora and fauna in Everglades National Park. Homestead, FL: Biological Resources Branch, South Florida Natural Resources Center, Everglades and Dry Tortugas National Parks.
- Hammond A, Pollegioni P, Persampieri T, North A, Minuz R, Trusso A, Bucci A, Kyrou K, Morianou I, Simoni A, Nolan T, Müller R, Crisanti A (2021) Gene-drive suppression of mosquito populations in large cages as a bridge between lab and field. Nature Communications 12(1): 4589. https://doi.org/10.1038/s41467-021-24790-6
- Hanslowe EB, Falk BG, Collier MAM, Josimovich JM, Rahill TA, Reed RN (2016) First record of invasive Burmese python oviposition and brooding inside an anthropogenic structure. Southeastern Naturalist (Steuben, ME) 15(sp8): 103–106. https://doi.org/10.1656/058.015.sp809
- Hanslowe EB, Duquesnel JG, Snow RW, Falk BG, Yackel Adams AA, Metzger III EF, Collier MAM, Reed RN (2018) Exotic predators may threaten another island ecosystem: A comprehensive assessment of python and boa reports from the Florida Keys. Management of Biological Invasions 9(3): 369–377. https://doi.org/10.3391/mbi.2018.9.3.18
- Harrison JB, Sunday JM, Rogers SM (2019) Predicting the fate of eDNA in the environment and implications for studying biodiversity. Proceedings. Biological Sciences 286(1915): 1409. https://doi.org/10.1098/rspb.2019.1409
- Hart KM, Schofield PJ, Gregoire DR (2012) Experimentally derived salinity tolerance of hatchling Burmese pythons (Python molurus bivittatus) from the Everglades, Florida (USA). Journal of Experimental Marine Biology and Ecology 413: 56–59. https://doi.org/10.1016/j.jembe.2011.11.021
- Hart KM, Cherkiss MS, Smith BJ, Mazzotti FJ, Fujisaki I, Snow RW, Dorcas ME (2015) Home range, habitat use, and movement patterns of non-native Burmese pythons in Everglades National Park, Florida, USA. Animal Biotelemetry 3(1): 1–13. https://doi.org/10.1186/s40317-015-0022-2
- Harvey RG, Perez L, Mazzotti FJ (2015) Not seeing is not believing: Volunteer beliefs about Burmese pythons in Florida and implications for public participation in invasive species removal. Journal of Environmental Planning and Management 59(5): 789–807. https://doi.org/10.1080/09640568.2015.1040489
- Harvey-Samuel T, Ant T, Alphey L (2017) Towards the genetic control of invasive species. Biological Invasions 19(6): 1683–1703. https://doi.org/10.1007/s10530-017-1384-6
- Hausfather Z, Drake HF, Abbott T, Schmidt GA (2020) Evaluating the performance of past climate model projections. Geophysical Research Letters 47: e2019GL085378. https://doi.org/10.1029/2019GL085378
- Heino M, Kaitala V (1999) Evolution of resource allocation between growth and reproduction in animals with indeterminate growth. Journal of Evolutionary Biology 12(3): 423–429. https://doi.org/10.1046/j.1420-9101.1999.00044.x
- Hendry AP, Castric V, Kinnison MT, Quinn TP, Stearns S (2004) The evolution of philopatry and dispersal. Evolution illuminated: salmon and their relatives. Oxford University Press, New York, NY, 52–90.
- Hengstebeck KC (2018) Burmese python use of gopher tortoise burrows in southwestern Florida. University of Florida.
- Hengstebeck KC, Romagosa CR (2020) A large trap for capturing large burrow-dwelling snakes. Herpetological Review, 51 pp.
- Hewitt J, Furxhi O, Renshaw CK, Driggers R (2021) Detection of Burmese pythons in the near-infrared versus visible band. Applied Optics 60(17): 5066–5073. https://doi.org/10.1364/AO.419320
- Hierink F, Bolon I, Durso AM, Ruiz de Castañeda R, Zambrana-Torrelio C, Eskew EA, Ray N (2020) Forty-four years of global trade in CITES-listed snakes: Trends and implications for conservation and public health. Biological Conservation 248: 108601. https://doi.org/10.1016/j.biocon.2020.108601
- Hillary RM, Bravington MV, Patterson TA, Grewe P, Bradford R, Feutry P, Gunasekera R, Peddemors V, Werry J, Francis MP, Duffy CAJ, Bruce BD (2018) Genetic relatedness reveals total population size of white sharks in eastern Australia and New Zealand. Scientific Reports 8(1): 2661. https://doi.org/10.1038/s41598-018-20593-w
- Holway D, Suarez A (1999) Animal behavior: An essential component of invasion biology. Trends in Ecology & Evolution 14(8): 328–330. https://doi.org/10.1016/S0169-5347(99)01636-5
- Hoon-Hanks LL, Ossiboff RJ, Bartolini P, Fogelson SB, Perry SM, Stöhr AC, Cross ST, Wellehan JFX, Jacobson ER, Dubovi EJ, Stenglein MD (2019) Longitudinal and cross-sectional sampling of serpentovirus (nidovirus) infection in captive snakes reveals high prevalence, persistent infection, and increased mortality in pythons and divergent serpentovirus infection in boas and colubrids. Frontiers in Veterinary Science 6: 1–17. https://doi.org/10.3389/fvets.2019.00338
- Hoover C (1998) The U.S. Role in the International Live Reptile Trade: Amazon Tree Boas to Zululand Dwarf Chameleons.
- Horak KE (2020) RNAi: Applications in Vertebrate Pest Management. Trends in Biotechnology 38(11): 1200–1202. https://doi.org/10.1016/j.tibtech.2020.05.001
- Hoyer IJ, Blosser EM, Acevedo C, Thompson AC, Reeves LE, Burkett-Cadena ND (2017) Mammal decline, linked to invasive Burmese python, shifts host use of vector mosquito towards reservoir hosts of a zoonotic disease. Biology Letters 13(10): 20170353. https://doi.org/10.1098/rsbl.2017.0353
- Humphrey J (2013) Trapping Method and Apparatus. U.S. Patent 8,407,931 B1.
- Hunter ME, Hart KM (2013) Rapid microsatellite marker development using next generation pyrosequencing to inform invasive Burmese python (Python molurus bivittatus) management. International Journal of Molecular Sciences 14(3): 4793–4804. https://doi.org/10.3390/ijms14034793
- Hunter ME, Oyler-McCance SJ, Dorazio RM, Fike JA, Smith BJ, Hunter CT, Reed RN, Hart KM (2015) Environmental DNA (eDNA) sampling improves occurrence and detection estimates of invasive Burmese pythons. PLoS ONE 10(4): 10. https://doi.org/10.1371/journal.pone.0121655
- Hunter ME, Johnson NA, Smith BJ, Davis MC, Butterfield JSS, Snow RW, Hart KM (2018) Cytonuclear discordance in the Florida Everglades invasive Burmese python (Python bivittatus) population reveals possible hybridization with the Indian python (P. molurus). Ecology and Evolution 8(17): 9034–9047. https://doi.org/10.1002/ece3.4423
- Hunter ME, Meigs-Friend G, Ferrante JA, Smith BJ, Hart KM (2019) Efficacy of eDNA as an early detection indicator for Burmese pythons in the ARM Loxahatchee National Wildlife Refuge in the greater Everglades ecosystem. Ecological Indicators 102: 617–622. https://doi.org/10.1016/j.ecolind.2019.02.058
- Hutchison VH, Dowling HG, Vinegar A (1966) Thermoregulation in a brooding female indian python, Python molurus bivittatus. Science 151(3711): 694–696. https://doi.org/10.1126/science.151.3711.694
- Imler RH (1945) Bullsnakes and their control on a Nebraska wildlife refuge. The Journal of Wildlife Management 9(4): 265–273. https://doi.org/10.2307/3796368
- Jacobs HJ, Auliya M, Böhme W, Auliya AVM, Jacobs HJ, Koch A, Wagner P (2009) Zur taxonomie des Dunklen Tigerpythons, Python molurus bivittatus Kuhl, 1820, speziell der Population von Sulawesi. Sauria 31: 5–16.
- Jacobson ER, Barker DG, Barker TM, Mauldin R, Avery ML, Engeman R, Secor S (2012) Environmental temperatures, physiology and behavior limit the range expansion of invasive Burmese pythons in southeastern USA. Integrative Zoology 7(3): 271–285. https://doi.org/10.1111/j.1749-4877.2012.00306.x
- Janousek WM, Hahn BA, Dreitz VJ (2019) Disentangling monitoring programs: Design, analysis, and application considerations. Ecological Applications 0(5): e01922. https://doi.org/10.1002/eap.1922
- Jeantet L, Planas-Bielsa V, Benhamou S, Geiger S, Martin J, Siegwalt F, Lelong P, Gresser J, Etienne D, Hiélard G, Arque A, Regis S, Lecerf N, Frouin C, Benhalilou A, Murgale C, Maillet T, Andreani L, Campistron G, Delvaux H, Guyon C, Richard S, Lefebvre F, Aubert N, Habold C, le Maho Y, Chevallier D (2020) Behavioural inference from signal processing using animal-borne multi-sensor loggers: A novel solution to extend the knowledge of sea turtle ecology. Royal Society Open Science 7(5): 7. https://doi.org/10.1098/rsos.200139
- Jenkins RKB, Brady LD, Huston K, Kauffmann JLD, Rabearivony J, Raveloson G, Rowcliffe JM (1999) The population status of chameleons within Ranomafana National Park, Madagascar, and recommendations for future monitoring. Oryx 33(1): 38–46. https://doi.org/10.1046/j.1365-3008.1999.00034.x
- Jerde CL, Mahon AR, Chadderton WL, Lodge DM (2011) “Sight-unseen” detection of rare aquatic species using environmental DNA. Conservation Letters 4(2): 150–157. https://doi.org/10.1111/j.1755-263X.2010.00158.x
- Jewell SD (2020) A century of injurious wildlife listing under the Lacey Act: A history. Management of Biological Invasions 11(3): 356–371. https://doi.org/10.3391/mbi.2020.11.3.01
- Johnson CR (1972) Thermoregulation in pythons-I. Effect of shelter, substrate type and posture on body temperature of the Australian carpet python, Morelia spilotes variegata. Comparative Biochemistry and Physiology. A. Comparative Physiology 43(2): 271–278. https://doi.org/10.1016/0300-9629(72)90185-5
- Johst K, Brandl R (1999) Natal versus breeding dispersal: Evolution in a model system. Evolutionary Ecology Research 1: 911–921.
- Jolley DB, Ditchkoff SS, Sparklin BD, Hanson LB, Mitchell MS, Grand JB (2010) Estimate of herpetofauna depredation by a population of wild pigs. Journal of Mammalogy 91(2): 519–524. https://doi.org/10.1644/09-MAMM-A-129.1
- Jordan F, Jelks HL, Kitchens WM (1997) Habitat structure and plant community composition in a northern Everglades wetland landscape. Wetlands 17(2): 275–283. https://doi.org/10.1007/BF03161415
- Josimovich JM, Currylow AF (2021) Hatchling growth experiment dataset from invasive Burmese pythons captured in 2015 in southern Florida: U.S. Geological Survey data release. https://doi.org/10.5066/P9WHSSJ6
- Josimovich JM, Falk BG, Grajal-Puche A, Hanslowe EB, Bartoszek IA, Reed RN, Currylow AF (2021) Clutch may predict growth of hatchling Burmese pythons better than food availability or sex. Biology Open 10(11): 1–10. https://doi.org/10.1242/bio.058739
- Kaiser BW, Osorio KJ, Enge KM, Engeman RM (2013) Tupinambis merinae (Argentine Giant Tegu); Pantherophis guttatus (Red Corn Snake) non-predatory killing. Herpetological Review 44: 329.
- Katz AD, Harper LR, Sternhagen EC, Pearce SE, Melder CA, Sperry JH, Davis MA (2021) Environmental DNA is effective in detecting the federally threatened Louisiana Pinesnake (Pituophis ruthveni). Environmental DNA 3(2): 409–425. https://doi.org/10.1002/edn3.126
- Kelehear C, Brown GP, Shine R (2013) Invasive parasites in multiple invasive hosts: The arrival of a new host revives a stalled prior parasite invasion. Oikos 122(9): 1317–1324. https://doi.org/10.1111/j.1600-0706.2013.00292.x
- Kelehear C, Spratt DM, O’Meally D, Shine R (2014) Pentastomids of wild snakes in the Australian tropics. International Journal for Parasitology. Parasites and Wildlife 3(1): 20–31. https://doi.org/10.1016/j.ijppaw.2013.12.003
- Kelly DW, Paterson RA, Townsend CR, Poulin R, Tompkins DM (2009) Parasite spillback: A neglected concept in invasion ecology? Ecology 90(8): 2047–2056. https://doi.org/10.1890/08-1085.1
- Kendall WL, Barker RJ, White GC, Lindberg MS, Langtimm CA, Peñaloza CL (2013) Combining dead recovery, auxiliary observations and robust design data to estimate demographic parameters from marked individuals. Methods in Ecology and Evolution 4(9): 828–835. https://doi.org/10.1111/2041-210X.12077
- Kéry M (2002) Inferring the absence of a species: A case study of snakes. The Journal of Wildlife Management 66(2): 330–338. https://doi.org/10.2307/3803165
- Kie JG, Matthiopoulos J, Fieberg J, Powell RA, Cagnacci F, Mitchell MS, Gaillard J-M, Moorcroft PR (2010) The home-range concept: Are traditional estimators still relevant with modern telemetry technology? Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences 365(1550): 2221–2231. https://doi.org/10.1098/rstb.2010.0093
- King MB, Duvall D (1990) Prairie rattlesnake seasonal migrations: Episodes of movement, vernal foraging and sex differences. Animal Behaviour 39(5): 924–935. https://doi.org/10.1016/S0003-3472(05)80957-1
- Klimley AP (2013) Why publish Animal Biotelemetry? Animal Biotelemetry 1(1): 2–4. https://doi.org/10.1186/2050-3385-1-1
- Kline E, Ripperger SP, Carter GG (2021) Habituation of common vampire bats to biologgers. Royal Society Open Science 8(12): 4–9. https://doi.org/10.1098/rsos.211249
- Kraus F (2009) Alien reptiles and amphibians: a scientific compendium and analysis. Springer Series in Invasion Biology, New York, NY, 369 pp. https://doi.org/10.1007/978-1-4020-8946-6
- Krishna SB (2002) Ophiophagus hannah (King Cobra). Diet. Herpetological Review 33: 141.
- Krysko KL, Nifong JC, Snow RW, Enge KM, Mazzotti FJ (2008) Reproduction of the Burmese python (Python molurus bivittatus) in southern Florida. Applied Herpetology 5(1): 93–95. https://doi.org/10.1163/157075408783489185
- Krysko KL, Hart KM, Smith BJ, Selby TH, Cherkiss MS, Coutu NT, Reichart RM, Nunez LP, Mazzotti FJ, Snow RW (2012) Record Length, Mass, and Clutch Size in the Nonindigenous Burmese Python, Python bivittatus Kuhl, 1820 (Squamata: Pythonidae), in Florida. Reptiles & Amphibians 19(4): 267–270. https://doi.org/10.17161/randa.v19i4.13923
- Krysko KL, Enge KM, Moler PE [Eds] (2019b) Amphibians and reptiles of Florida. University of Florida Press, Gainesville, 728 pp.
- Krysko KL, Reed RN, Rochford MR, Nunez LP, Enge KM (2019a) Python bivittatus Kuhl, 1820, Burmese python. Species account. The Amphibians and Reptiles of Florida. University of Florida Press, Gainesville, 458–460.
- Kucherenko A, Herman JE, Everham III EM, Urakawa H (2018) Terrestrial snake environmental DNA accumulation and degradation dynamics and its environmental application. Herpetologica 74(1): 38–49. https://doi.org/10.1655/Herpetologica-D-16-00088
- Lantz SM, Gawlik DE, Cook MI (2010) The effects of water depth and submerged aquatic vegetation on the selection of foraging habitat and foraging succes of wading birds. The Condor 112(3): 460–469. https://doi.org/10.1525/cond.2010.090167
- Lardner B, Yackel Adams AA, Savidge JA, Rodda GH, Reed RN, Clark CS (2013) Effectiveness of bait tubes for brown treesnake control on Guam. Wildlife Society Bulletin 37: 664–673. https://doi.org/10.1002/wsb.297
- Lebreton JD (2005) Dynamical and statistical models for exploited populations. Australian & New Zealand Journal of Statistics 47(1): 49–63. https://doi.org/10.1111/j.1467-842X.2005.00371.x
- Lebreton JD, Burnham KP, Clobert J, Anderson DR (1992) Modeling survival and testing biological hypotheses using marked animals: A unified approach with case studies. Ecological Monographs 62(1): 67–118. https://doi.org/10.2307/2937171
- Light S, Dineen J (1994) Water control in the Everglades: a historical perspective. Everglades: the ecosystem and its restoration. CRC Press, Boca Raton, FL, 47–84.
- Lillywhite HB, De Delva P, Noonan BP (2002) Patterns of gut passage time and the chronic retention of fecal mass in viperid snakes. Biology of the Vipers. Eagle Mountain Pub Lc, 497–506.
- Link WA, Converse SJ, Yackel Adams AA, Hostetter NJ (2018) Analysis of population change and movement using robust design removal data. Journal of Agricultural Biological & Environmental Statistics 23(4): 463–477. https://doi.org/10.1007/s13253-018-0335-8
- Lodge TE (2010) The Everglades Handbook: Understanding the ecosystem. 3rd Edn. CRC Press, 422 pp.
- Lorch JM, Lankton J, Werner K, Falendysz EA, McCurley K, Blehert DS (2015) Experimental infection of snakes with Ophidiomyces ophiodiicola causes pathological changes that typify snake fungal disease. mBio 6(6): 6. https://doi.org/10.1128/mBio.01534-15
- MacKenzie DI, Nichols JD, Lachman GB, Droege S, Royle AA, Langtimm CA (2002) Estimating site occupancy rates when detection probabilities are less than one. Ecology 83(8): 2248–2255. https://doi.org/10.1890/0012-9658(2002)083[2248:ESORWD]2.0.CO;2
- MacKenzie DI, Nichols JD, Royle JA, Pollock KH, Bailey LL, Hines JE (2017) Occupancy Estimation and Modeling: Inferring Patterns and Dynamics of Species Occurrence: Second Edition. Page Occupancy Estimation and Modeling: Inferring Patterns and Dynamics of Species Occurrence: 2nd Edn. Elsevier, 648 pp. https://doi.org/10.1016/B978-0-12-407197-1.00019-3
- Madsen T, Shine R (1993) Phenotypic plasticity in body sizes and sexual size dimorphism in European grass snakes. Evolution; International Journal of Organic Evolution 47: 321–325. https://doi.org/10.1111/j.1558-5646.1993.tb01222.x
- Madsen T, Shine R (2000) Silver spoons and snake body sizes: Prey availability early in life influences long-term growth rates of free-ranging pythons. Journal of Animal Ecology 69(6): 952–958. https://doi.org/10.1046/j.1365-2656.2000.00477.x
- Madsen T, Ujvari B, Shine R, Olsson M (2006) Rain, rats and pythons: Climate-driven population dynamics of predators and prey in tropical Australia. Austral Ecology 31(1): 30–37. https://doi.org/10.1111/j.1442-9993.2006.01540.x
- Maehr DS, Brady JR, Maehr DS, Brady JR (1986) Food habits of bobcats in Florida. Journal of Mammalogy 67(1): 133–138. https://doi.org/10.2307/1381009
- Mason RT, Parker MR (2010) Social behavior and pheromonal communication in reptiles. Journal of Comparative Physiology. A, Neuroethology, Sensory, Neural, and Behavioral Physiology 196(10): 729–749. https://doi.org/10.1007/s00359-010-0551-3
- Mason RT, Fales HM, Jones TH, Pannell LK, Chinn W, Crews D (1989) Sex pheromones in snakes. Science 245(4915): 290–293. https://doi.org/10.1126/science.2749261
- Matechou E, McCrea RS, Morgan BJT, Nash DJ, Griffiths RA (2016) Open models for removal data. The Annals of Applied Statistics 10(3): 1572–1589. https://doi.org/10.1214/16-AOAS949
- Mauldin RE, Savarie PJ (2010) Acetaminophen as an oral toxicant for Nile monitor lizards (Varanus niloticus) and Burmese pythons (Python molurus bivittatus). Wildlife Research 37(3): 215–222. https://doi.org/10.1071/WR08168
- Mazzotti FJ, Cherkiss MS, Hart KM, Snow RW, Rochford MR, Dorcas ME, Reed RN (2011) Cold-induced mortality of invasive Burmese pythons in south Florida. Biological Invasions 13(1): 143–151. https://doi.org/10.1007/s10530-010-9797-5
- Mazzotti FJ, Mceachern M, Rochford M, Reed RN, Eckles JK, Vinci J, Edwards J, Wasilewski J (2015) Tupinambis merianae as nest predators of crocodilians and turtles in Florida, USA. Biological Invasions 17(1): 47–50. https://doi.org/10.1007/s10530-014-0730-1
- Mazzotti FJ, Rochford M, Vinci J, Jeffery BM, Eckles JK, Dove C, Sommers KP (2016) Implications of the 2013 Python Challenge for Ecology and Management of Python molurus bivittatus (Burmese Python) in Florida. Southeastern Naturalist (Steuben, ME) 15(sp8): 63–74. https://doi.org/10.1656/058.015.sp807
- Mazzotti FJ, Nestler JH, Cole JM, Closius C, Kern WH, Rochford MR, Suarez E, Brubaker R, Platt SG, Rainwater T, Ketterlin JK (2020) Diet of Nile monitors (Varanus niloticus) Removed from Palm Beach and Broward Counties, Florida, USA. Journal of Herpetology 54(2): 189–195. https://doi.org/10.1670/18-115
- McCaffrey KR, Miller MA, Balaguera-Reina SA, Mazzotti FJ (2022) 2022 Final Report to the Florida Fish and Wildlife Conservation Commission (FWC): Assistance in Data Management and Analysis for the Python Action Team Removing Invasive Constrictors (PATRIC) Program. Contract Number 13416–A3051.
- McCleery RA, Sovie A, Reed RN, Cunningham MW, Hunter ME, Hart KM (2015) Marsh rabbit mortalities tie pythons to the precipitous decline of mammals in the Everglades. Proceedings. Biological Sciences 282(1805): 20150120. https://doi.org/10.1098/rspb.2015.0120
- McClintock BT, London JM, Cameron MF, Boveng PL (2017) Bridging the gaps in animal movement: Hidden behaviors and ecological relationships revealed by integrated data streams. Ecosphere 8(3): 8. https://doi.org/10.1002/ecs2.1751
- McCollister MF, Josimovich JM, Fitzgerald AL, Jansen DK, Currylow AF (2021) Native mammalian predators can depredate adult Burmese pythons in Florida. Southeastern Naturalist (Steuben, ME) 20(55–N): 59. https://doi.org/10.1656/058.020.0205
- McDiarmid RW, Campbell JA, Toure T (1999) Snake species of the world: A taxonomic and geographic reference. Volume 1. Herpetologists’ League.
- Means DB (2017) Diamonds in the rough: Natural history of the Eastern Diamondback rattlesnake. Tall Timbers Press.
- Meshaka Jr WE, Loftus WF, Steiner T (2000) The herpetofauna of everglades National Park. Florida Academy of Sciences, Inc. 63: 84–103.
- Metcalf MF, Marsh A, Brosse W, Herman JE (2019) Crotalus adamanteus (Eastern Diamondback Rattlesnake) endoparasite. Herpetological Review 50: 389.
- Metzger CJ (2013) Python molurus bivittatus (Burmese Python). Habitat use/occurance within Gopherus polyphemus burrows. Herpetological Review 44: 333–334.
- Miller MA, Kinsella JM, Snow RW, Hayes MM, Falk BG, Reed RN, Mazzotti FJ, Guyer C, Romagosa CM (2018) Parasite spillover: Indirect effects of invasive Burmese pythons. Ecology and Evolution 8(2): 830–840. https://doi.org/10.1002/ece3.3557
- Miller MA, Kinsella JM, Snow RW, Falk BG, Reed RN, Goetz SM, Mazzotti FJ, Guyer C, Romagosa CM (2020) Highly competent native snake hosts extend the range of an introduced parasite beyond its invasive Burmese python host. Ecosphere: 11(6): e03153. https://doi.org/10.1002/ecs2.3153
- Miralles L, Dopico E, Devlo-Delva F, Garcia-Vazquez E (2016) Controlling populations of invasive pygmy mussel (Xenostrobus securis) through citizen science and environmental DNA. Marine Pollution Bulletin 110(1): 127–132. https://doi.org/10.1016/j.marpolbul.2016.06.072
- Mitsch WJ, Hernandez ME (2013) Landscape and climate change threats to wetlands of North and Central America. Aquatic Sciences 75(1): 133–149. https://doi.org/10.1007/s00027-012-0262-7
- Moe SJ, Stenseth NC, Smith RH (2002) Density-dependent compensation in blowfly populations give indirectly positive effects of a toxicant. Ecology 83(6): 1597–1603. https://doi.org/10.1890/0012-9658(2002)083[1597:DDCIBP]2.0.CO;2
- Montes E, Pleguezuelos JM, Kraus F (2021) Rapid endangerment of the lizard Podarcis pityusensis by an invasive snake demands an immediate conservation response. Amphibian & Reptile Conservation 15: 238–243.
- Moran AP (1951) A mathematical theory of animal trapping. Biometrika 38(3–4): 307–311. https://doi.org/10.1093/biomet/38.3-4.307
- Mortensen HS, Dupont YL, Olesen JM (2008) A snake in paradise: Disturbance of plant reproduction following extirpation of bird flower-visitors on Guam. Biological Conservation 141(8): 2146–2154. https://doi.org/10.1016/j.biocon.2008.06.014
- Moyer GR, Díaz-Ferguson E, Hill JE, Shea C (2014) Assessing environmental DNA detection in controlled lentic systems. PLoS ONE 9(7): 9. https://doi.org/10.1371/journal.pone.0103767
- Murray DL (2006) On improving telemetry-based survival estimation. The Journal of Wildlife Management 70(6): 1530–1543. https://doi.org/10.2193/0022-541X(2006)70[1530:OITSE]2.0.CO;2
- Mushinsky HR, Miller DE (1993) Predation on Water Snakes: Ontogenetic and interspecific considerations. Copeia 1993(3): 660–665. https://doi.org/10.2307/1447227
- Mutascio HE, Pittman SE, Zollner PA (2017) Investigating movement behavior of invasive Burmese pythons on a shy-bold continuum using individual-based modeling. Perspectives in Ecology and Conservation 15(1): 25–31. https://doi.org/10.1016/j.pecon.2017.02.004
- Mutascio HE, Pittman SE, Zollner PA, D’Acunto LE (2018) Modeling relative habitat suitability of southern Florida for invasive Burmese pythons (Python molurus bivittatus). Landscape Ecology 33(2): 257–274. https://doi.org/10.1007/s10980-017-0597-5
- Nafus MG, Mazzotti FJ, Reed RN (2020) Estimating detection probability for Burmese pythons with few detections and zero recaptures. Journal of Herpetology 54(1): 24–30. https://doi.org/10.1670/18-154
- Nams VO (2006) Detecting oriented movement of animals. Animal Behaviour 72(5): 1197–1203. https://doi.org/10.1016/j.anbehav.2006.04.005
- NAS (2016) National Academies of Sciences. Gene drives on the horizon: Advancing science, navigating uncertainty, and aligning research with public values. The National Academies Press, Washington, D.C.
- Nathan R, Getz WM, Revilla E, Holyoak M, Kadmon R, Saltz D, Smouse PE (2008) A movement ecology paradigm for unifying organismal movement research. Proceedings of the National Academy of Sciences of the United States of America 105(49): 19052–19059. https://doi.org/10.1073/pnas.0800375105
- Newcomb KC, Davis JB, Kaminski RM, Gray MJ (2016) Winter survival of female American black ducks in Tennessee, USA. The Condor 118(1): 33–45. https://doi.org/10.1650/CONDOR-15-74.1
- Newsome SD, Martinez del Rio C, Bearhop S, Phillips DL (2007) A niche for isotopic ecology. Frontiers in Ecology and the Environment 5(8): 429–436.
- Nicholson AJ (1957) The self-adjustment of populations to change. Cold Spring Harbor Symposia on Quantitative Biology 22(0): 153–173. https://doi.org/10.1101/SQB.1957.022.01.017
- O’Dea MA, Jackson B, Jackson C, Xavier P, Warren K (2016) Discovery and partial genomic characterisation of a novel nidovirus associated with respiratory disease in wild shingleback lizards (Tiliqua rugosa). PLoS ONE 11(11): e0165209. https://doi.org/10.1371/journal.pone.0165209
- O’Shea M (2007) Boas and pythons of the world. Princeton University Press, Princeton, NJ, 160 pp.
- Orem W, Gilmour C, Axelrad D, Krabbenhoft D, Scheidt D, Kalla P, McCormick P, Gabriel M, Aiken G (2011) Sulfur in the South Florida ecosystem: Distribution, sources, biogeochemistry, impacts, and management for restoration. Critical Reviews in Environmental Science and Technology 41(sup1): 249–288. https://doi.org/10.1080/10643389.2010.531201
- Orzechowski SCM, Frederick PC, Dorazio RM, Hunter ME (2019a) Environmental DNA sampling reveals high occupancy rates of invasive Burmese pythons at wading bird breeding aggregations in the central Everglades. PLoS ONE 14(4): 1–18. https://doi.org/10.1371/journal.pone.0213943
- Orzechowski SCM, Romagosa CM, Frederick PC (2019b) Invasive Burmese pythons (Python bivittatus) are novel nest predators in wading bird colonies of the Florida Everglades. Biological Invasions 21(7): 2333–2344. https://doi.org/10.1007/s10530-019-01979-x
- Otis DL, Burnham KP, White GC, Anderson DR (1978) Statistical inference from capture data on closed animal populations. Wildlife Monographs, 3–135.
- Oye KA, Esvelt K, Appleton E, Catteruccia F, Church G, Kuiken T, Lightfoot SB-Y, McNamara J, Smidler A, Collins JP (2014) Regulating gene drives. Science 345(6197): 626–628. https://doi.org/10.1126/science.1254287
- Pace ML, Cole JJ, Carpenter SR, Kitchell JF (1999) Trophic cascades revealed in diverse ecosystems. Trends in Ecology & Evolution 14(12): 483–488. https://doi.org/10.1016/S0169-5347(99)01723-1
- Palmisano JN, Bockoven C, McPherson SM, Ossiboff RJ, Walden HDS, Farrell TM (2022) Exposure experiments indicate some common Florida anurans and lizards may serve as intermediate hosts for the invasive pentastome parasite, Raillietiella orientalis. Journal of Herpetology 56(3). https://doi.org/10.1670/21-061
- Pardini EA, Drake JM, Chase JM, Knight TM (2009) Complex population dynamics and control of the invasive biennial Alliaria petiolata (garlic mustard). Ecological Applications 19(2): 387–397. https://doi.org/10.1890/08-0845.1
- Parker MR, Mason RT (2011) Pheromones in Snakes: History, Patterns and Future Research Directions. Reproductive Biology and Phylogeny of Snakes, 551–572. https://doi.org/10.1201/b10879-14
- Parker MR, Mason RT (2012) How to make a sexy snake: Estrogen activation of female sex pheromone in male red-sided garter snakes. The Journal of Experimental Biology 215(5): 723–730. https://doi.org/10.1242/jeb.064923
- Parker MR, Mason RT (2014) A novel mechanism regulating a sexual signal: The testosterone-based inhibition of female sex pheromone expression in garter snakes. Hormones and Behavior 66(3): 509–516. https://doi.org/10.1016/j.yhbeh.2014.07.007
- Parker WS, Plummer MV (1987) Population Ecology. Eds. Snakes: Ecology and Evolutionary Biology. The Blackburn Press, Caldwell, New Jersey, 253‐301.
- Parker MR, Patel SM, Zachry JE, Kimball BA (2018) Feminization of Male Brown Treesnake Methyl Ketone Expression via Steroid Hormone Manipulation. Journal of Chemical Ecology 44(2): 189–197. https://doi.org/10.1007/s10886-018-0935-3
- Parker MR, Currylow AF, Tillman EA, Robinson IM, Josimovich CJ, Bukovich JM, Nazarian LA, Nafus MG, Kluever BM, Yackel Adams AA (2021) Using enclosed Y-Mazes to assess chemosensory behavior in reptiles. Journal of Visualized Experiments Apr 7: e61858. https://doi.org/10.3791/61858
- Parrish K, Kirkland PD, Skerratt LF, Ariel E (2021) Nidoviruses in reptiles: A review. Frontiers in Veterinary Science (8): 733404. https://doi.org/10.3389/fvets.2021.733404
- Pasko S, Goldberg J (2014) Review of harvest incentives to control invasive species. Management of Biological Invasions 5(3): 263–277. https://doi.org/10.3391/mbi.2014.5.3.10
- Pavlacky Jr DC, Blakesley JA, White GC, Hanni DJ, Lukacs PM (2012) Hierarchical multi-scale occupancy estimation for monitoring wildlife populations. The Journal of Wildlife Management 76(1): 154–162. https://doi.org/10.1002/jwmg.245
- Pearson D, Shine R, Williams A (2005) Spatial ecology of a threatened python (Morelia spilota imbricata) and the effects of anthropogenic habitat change. Austral Ecology 30(3): 261–274. https://doi.org/10.1111/j.1442-9993.2005.01462.x
- Perry G, Morton JM (1999) Regeneration rates of the woody vegetation of guam’s northwest field following major disturbance: Land use patterns, feral ungulates, and cascading effects of the brown treesnake. Micronesica 32: 125–142.
- Phillips SJ, Anderson RP, Schapire RE (2006) Maximum entropy modeling of species geographic distributions. Ecological Modelling 190(3–4): 231–259. https://doi.org/10.1016/j.ecolmodel.2005.03.026
- Piaggio AJ, Engeman RM, Hopken MW, Humphrey JS, Keacher KL, Bruce WE, Avery ML (2014) Detecting an elusive invasive species: A diagnostic PCR to detect Burmese python in Florida waters and an assessment of persistence of environmental DNA. Molecular Ecology Resources 14(2): 374–380. https://doi.org/10.1111/1755-0998.12180
- Picardi S, Frederick PC, Borkhataria RR, Basille M (2020) Partial migration in a subtropical wading bird in the southeastern United States. Ecosphere 11(2): e03054. https://doi.org/10.1002/ecs2.3054
- Pifer EK, Hart KM, Rice KG, Mazzotti FJ (2009) Final Report for CESU Cooperative Agreement H5000060106 and Task Agreement J2117062273. Small and Medium Sized Mammal Inventory for Everglades National Park and Big Cypress National Preserve.
- Piquet JC, López-Darias M (2021) Invasive snake causes massive reduction of all endemic herpetofauna on Gran Canaria. Proceedings of the Royal Society B: Biological Sciences, 288 pp. https://doi.org/10.1098/rspb.2021.1939
- Pittman SE, Bartoszek IA (2021) Initial dispersal behavior and survival of non-native juvenile Burmese pythons (Python bivittatus) in South Florida. BMC Zoology 6(1): 1–13. https://doi.org/10.1186/s40850-021-00098-2
- Pittman SE, Hart KM, Cherkiss MS, Snow RW, Fujisaki I, Smith BJ, Mazzotti FJ, Dorcas ME (2014) Homing of invasive Burmese pythons in South Florida: Evidence for map and compass senses in snakes. Biology Letters 10(3): 10. https://doi.org/10.1098/rsbl.2014.0040
- Poghosyan A, Golkar A (2017) CubeSat evolution: Analyzing CubeSat capabilities for conducting science missions. Progress in Aerospace Sciences 88: 59–83. https://doi.org/10.1016/j.paerosci.2016.11.002
- Pollock KH (1976) Building models of capture-recapture experiments. Journal of the Royal Statistical Society. Series A (General) 25: 253–259. https://doi.org/10.2307/2988083
- Pollock KH, Nichols JD, Brownie C, Hines JE (1990) Statistical inference for capture-recapture experiments. Wildlife Monographs, 3–97.
- Pope CH (1961) The giant snakes: the natural history of the Boa constrictor, the anaconda, and the largest pythons, including comparative facts about other snakes and basic information on reptiles in general. Random House Inc.
- Pough FH (1980) The advantages of ectothermy for tetrapods. American Naturalist 115(1): 92–112. https://doi.org/10.1086/283547
- Powell RA, Mitchell MS (2012) What is a home range? Journal of Mammalogy 93(4): 948–958. https://doi.org/10.1644/11-MAMM-S-177.1
- Pyron RA, Burbrink FT, Guiher TJ (2008) Claims of potential expansion throughout the U.S. by invasive python species are contradicted by ecological niche models. PLoS ONE 3(8): e2931. https://doi.org/10.1371/journal.pone.0002931
- Rahman SC (2013) Python molurus bivittatus (Burmese python). Reproduction/Nesting. Herpetological Review 44: 334.
- Rajesh NV, Rajesh KD, Jayathangaraj MG, Raman M, Sridhar R (2015) Parasitic fauna of captive snakes in Tamilnadu, India. Asian Pacific Journal of Tropical Disease 5(7): 547–551. https://doi.org/10.1016/S2222-1808(15)60834-9
- Randhawa M, Seo IS, Liebel F, Southall MD, Kollias N, Ruvolo E (2015) Visible light induces melanogenesis in human skin through a photoadaptive response. PLoS ONE 10(6): 1–14. https://doi.org/10.1371/journal.pone.0130949
- Rasys AM, Park S, Ball RE, Alcala AJ, Lauderdale JD, Menke DB (2019) CRISPR-Cas9 Gene Editing in Lizards through Microinjection of Unfertilized Oocytes. Cell Reports 28(9): 2288–2292. https://doi.org/10.1016/j.celrep.2019.07.089
- Réale D, Reader SM, Sol D, McDougall PT, Dingemanse NJ (2007) Integrating animal temperament within ecology and evolution. Biological Reviews of the Cambridge Philosophical Society 82(2): 291–318. https://doi.org/10.1111/j.1469-185X.2007.00010.x
- Redford KH, Brooks TM, Macfarlane NBW, Adams JS [Eds] (2019) Genetic frontiers for conservation: an assessment of synthetic biology and biodiversity conservation. Technical assessment. Page Genetic frontiers for conservation: an assessment of synthetic biology and biodiversity conservation: technical assessment. IUCN, International Union for Conservation of Nature, 1–184. https://doi.org/10.2305/IUCN.CH.2019.05.en
- Reed RN, Hart KM, Rodda GH, Mazzotti FJ, Snow RW, Cherkiss M, Rozar R, Goetz S (2011) A field test of attractant traps for invasive Burmese pythons (Python molurus bivittatus) in southern Florida. Wildlife Research 38(2): 114–121. https://doi.org/10.1071/WR10202
- Reed RN, Kraus F (2010) Invasive reptiles and amphibians: Global perspectives and local solutions. Animal Conservation 13: 3–4. https://doi.org/10.1111/j.1469-1795.2010.00409.x
- Reed RN, Rodda GH (2009) Giant constrictors: Biological and management profiles and an establishment risk assessment for nine large species of pythons, anacondas, and the Boa constrictor. https://doi.org/10.3133/ofr20091202
- Reed RN, Shine R, Shetty S (2002) Sea kraits (Squamata: Laticauda spp.) as a useful bioassay for assessing local diversity of eels (Muraenidae, Congridae) in the Western Pacific Ocean. Copeia 2002(4): 1098–1101. https://doi.org/10.1643/0045-8511(2002)002[1098:SKSLSA]2.0.CO;2
- Reed RN, Snow RW (2014) Assessing risks to humans from invasive burmese pythons in Everglades National Park, Florida, USA. Wildlife Society Bulletin 38(2): 366–369. https://doi.org/10.1002/wsb.413
- Rees HC, Maddison BC, Middleditch DJ, Patmore JRM, Gough KC (2014) The detection of aquatic animal species using environmental DNA-a review of eDNA as a survey tool in ecology. Journal of Applied Ecology 51(5): 1450–1459. https://doi.org/10.1111/1365-2664.12306
- Reeves LE, Krysko KL, Avery ML, Gillett-Kaufman JL, Kawahara AY, Connelly CR, Kaufman PE (2018) Interactions between the invasive Burmese python, Python bivittatus Kuhl, and the local mosquito community in Florida, USA. PLoS ONE 13(1): e0190633. https://doi.org/10.1371/journal.pone.0190633
- Reichert BE, Sovie AR, Udell BJ, Hart KM, Borkhataria RR, Bonneau M, Reed R, McCleery R (2017) Urbanization may limit impacts of an invasive predator on native mammal diversity. Diversity & Distributions 23(4): 355–367. https://doi.org/10.1111/ddi.12531
- Reilly SB, Gottscho AD, Garwood JM, Jennings WB (2010) Phylogenetic analysis of Common Garter Snake (Thamnophis sirtalis) stomach contents detects cryptic range of a secretive salamander (Ensatina eschscholtzii oregonensis). Herpetological Conservation and Biology 5: 395–402.
- Reynolds GR, Niemiller ML, Revell LJ (2014) Toward a Tree-of-Life for the boas and pythons: Multilocus species-level phylogeny with unprecedented taxon sampling. Molecular Phylogenetics and Evolution 71: 201–213. https://doi.org/10.1016/j.ympev.2013.11.011
- Richard SA, Tillman EA, Humphrey JS, Avery ML, Parker MR (2019) Male Burmese pythons follow female scent trails and show sex-specific behaviors. Integrative Zoology 14(5): 460–469. https://doi.org/10.1111/1749-4877.12376
- Ripperger SP, Carter GG, Page Supervision RA, Duda N, Koelpin A, Weigel R, Hartmann M, Nowak T, Thielecke J, Schadhauser M, Robert J, Herbst S, Meyer-Wegener K, Wägemann P, Preikschat WS, Cassens B, Kapitza R, Dressler F, Mayer F (2020) Thinking small: Next-generation sensor networks close the size gap in vertebrate biologging. PLoS Biology 18(4): 1–25. https://doi.org/10.1371/journal.pbio.3000655
- Rivas JA, Burghardt GM (2005) Snake mating systems, behavior, and evolution: The revisionary implications of recent findings. Journal of Comparative Psychology (Washington, D.C. ) 119(4): 447–454. https://doi.org/10.1037/0735-7036.119.4.447
- Rochford M, Brien ML, Carrigan J, Snow RW, Mazzotti FJ (2010a) Python molurus bivittatus (Burmese python). Clutch size. Herpetological Review 41: 97.
- Rochford M, Krysko KL, Nifong J, Wilkins L, Snow R, Cherkiss MS (2010b) Python molurus bivittatus (Burmese Python). Diet. Herpetological Review 41: 97.
- Rodda GH, Campbell EW (2002) Distance sampling of forest snakes and lizards. Herpetological Review 33: 271–274.
- Rodda GH, Fritts TH, Clark CS, Gotte SW, Chiszar D (1999) A state-of-the-art trap for the brown treesnake. Problem snake management: The habu and the brown treesnake. Comstock Publishing Associates, Ithaca, NY, 268–284. https://doi.org/10.7591/9781501737688
- Rodda GH, Savidge JA, Tyrrell CL, Christy MT, Ellingson AR (2007) Size Bias in Visual Searches and Trapping of Brown Treesnakes on Guam. The Journal of Wildlife Management 71(2): 656–661. https://doi.org/10.2193/2005-742
- Rodda GH, Jarnevich CS, Reed RN (2009) What parts of the US mainland are climatically suitable for invasive alien pythons spreading from Everglades National Park? Biological Invasions 11(2): 241–252. https://doi.org/10.1007/s10530-008-9228-z
- Rodda GH, Jarnevich CS, Reed RN (2011) Challenges in identifying sites climatically matched to the native ranges of animal invaders. PLoS ONE 6(2): e14670. https://doi.org/10.1371/journal.pone.0014670
- Rogers HS, Buhle ER, Hille Ris Lambers J, Fricke EC, Miller RH, Tewksbury JJ (2017) Effects of an invasive predator cascade to plants via mutualism disruption. Nature Communications 8(1): 6–13. https://doi.org/10.1038/ncomms14557
- Rogers H, Hille Ris Lambers J, Miller R, Tewksbury JJ (2012) “Natural experiment” Demonstrates top-down control of spiders by birds on a landscape level. PLoS ONE 7(9): e43446. https://doi.org/10.1371/journal.pone.0043446
- Romagosa CM (2014) Patterns of live vertebrate importation into the United States: analysis of an invasion pathway. In: Keller RP, et al. (Eds) Invasive species in a globalized world: ecological, social, and legal perspectives on policy. University of Chicago Press, Illinois, 115-146.
- Romagosa CM, Steury T, Miller MA, Guyer C, Rogers B, Angle TC, Gillette RL (2011) Assessment of detection dogs as a potential tool for python detection efforts. Final report submitted to the National Park service-Everglades National Park (Contract H500002A271) and South Florida Water Management District (Contract 4500059338).
- Romagosa CM, Mazzotti FJ, Snow RW, Bartoszek I, Dove CJ, Diego Juárez-Sánchez A, Suarez E, Rochford MR, Krysko KL, Cherkiss MS, Falk BG, Josimovich JM, Yackel Adams AA, Currylow AF, Hart KM, Reed RN (2022) Diet of invasive Burmese Pythons (Python molurus bivittatus) in southern Florida, 1995–2020. US Geological Survey Data Release. https://doi.org/10.5066/P9A0V89V
- Royle JA, Fuller AK, Sutherland C (2018) Unifying population and landscape ecology with spatial capture-recapture. Ecography 41(3): 444–456. https://doi.org/10.1111/ecog.03170
- Ruiz P, Laplanche C (2010) A hierarchical model to estimate the abundance and biomass of salmonids by using removal sampling and biometric data from multiple locations. Canadian Journal of Fisheries and Aquatic Sciences 67(12): 2032–2044. https://doi.org/10.1139/F10-123
- Rumbold DG, Bartoszek IA (2019) Mercury concentrations in invasive Burmese pythons (Python bivittatus) of Southwest Florida. Bulletin of Environmental Contamination and Toxicology 103(4): 533–537. https://doi.org/10.1007/s00128-019-02670-6
- Ruzzante DE, McCracken GR, Førland B, MacMillan J, Notte D, Buhariwalla C, Mills Flemming J, Skaug H (2019) Validation of close-kin mark-recapture (CKMR) methods for estimating population abundance. Methods in Ecology and Evolution 10(9): 1445–1453. https://doi.org/10.1111/2041-210X.13243
- Savarie PJ, Shivik JA, White GC, Hurley JC, Clark L (2001) Use of acetaminophen for large-scale control of brown treesnakes. The Journal of Wildlife Management 65(2): 356. https://doi.org/10.2307/3802916
- Savidge JA (1987) Extinction of an Island Forest Avifauna by an Introduced Snake. Ecology 68(3): 660–668. https://doi.org/10.2307/1938471
- Schaub M, Abadi F (2011) Integrated population models: A novel analysis framework for deeper insights into population dynamics. Journal of Ornithology 152(S1): S227–S237. https://doi.org/10.1007/s10336-010-0632-7
- Schleip WD, O’Shea M (2010) Annotated checklist of the recent and extinct pythons (Serpentes, Pythonidae), with notes on nomenclature, taxonomy, and distribution. ZooKeys 66: 29–79. https://doi.org/10.3897/zookeys.66.683
- Schmidt KP (1932) Stomach contents of some American coral snakes, with the description of a new species of Geophis. Copeia 1932(1): 6–9. https://doi.org/10.2307/1437020
- Schmidt TW (1979) 1979 Ecological study of fishes and the water quality characteristics of Florida Bay, Everglades National Park, Florida. NOAA/National Ocean Service/National Centers for Coastal Ocean Science.
- Schmidt BR, Kéry M, Ursenbacher S, Hyman OJ, Collins JP (2013) Site occupancy models in the analysis of environmental DNA presence/absence surveys: A case study of an emerging amphibian pathogen. Methods in Ecology and Evolution 4(7): 646–653. https://doi.org/10.1111/2041-210X.12052
- Schwarz CJ, Seber GAF (1999) Estimating animal abundance: Review III. Statistical Science 14(4): 427–456. https://doi.org/10.1214/ss/1009212521
- Seber GAF (1986) A Review of Estimating Animal Abundance. Biometrics 42(2): 267–292. https://doi.org/10.2307/2531049
- Secor SM (2008) Digestive physiology of the Burmese python: Broad regulation of integrated performance. The Journal of Experimental Biology 211(24): 3767–3774. https://doi.org/10.1242/jeb.023754
- Secor SM, Diamond J (1995) Adaptive responses to feeding in Burmese pythons: Pay before pumping. The Journal of Experimental Biology 198(6): 1313–1325. https://doi.org/10.1242/jeb.198.6.1313
- Secor SM, Diamond JM (2000) Evolution of regulatory responses to feeding in snakes. Physiological and Biochemical Zoology 73(2): 123–141. https://doi.org/10.1086/316734
- Séret B, Brischoux F, Bonnet X, Shine R (2008) First record of Cirrimaxilla formosa (Muraenidae) from New Caledonia, found in sea snake stomach contents. Cybium 32: 191–192.
- Sewell D, Guillera-Arroita G, Griffiths RA, Beebee TJC (2012) When is a species declining? Optimizing survey effort to detect population changes in reptiles. PLoS ONE 7(8): e43387. https://doi.org/10.1371/journal.pone.0043387
- SFWMD (2022) South Florida Water Management District. Python Elimination Program.
- Shackleton RT, Adriaens T, Brundu G, Dehnen-Schmutz K, Estévez RA, Fried J, Larson BMH, Liu S, Marchante E, Marchante H, Moshobane MC, Novoa A, Reed M, Richardson DM (2019) Stakeholder engagement in the study and management of invasive alien species. Journal of Environmental Management 229: 88–101. https://doi.org/10.1016/j.jenvman.2018.04.044
- Shine R (2005) Life-history evolution in reptiles. Annual Review of Ecology, Evolution, and Systematics 36(1): 23–46. https://doi.org/10.1146/annurev.ecolsys.36.102003.152631
- Shirk PL, Linden DW, Patrick DA, Howell KM, Harper EB, Vonesh JR (2014) Impact of habitat alteration on endemic Afromontane chameleons: Evidence for historical population declines using hierarchical spatial modelling. Diversity & Distributions 20(10): 1186–1199. https://doi.org/10.1111/ddi.12239
- Siers SR, Yackel Adams AA, Reed RN (2018) Behavioral differences following ingestion of large meals and consequences for management of a harmful invasive snake: A field experiment. Ecology and Evolution 8(20): 10075–10093. https://doi.org/10.1002/ece3.4480
- Siers SR, Pitt W, Eisemann J, Clark L, Shiels AB (2019) In situ evaluation of an automated aerial bait delivery system for landscape-scale control of invasive brown treesnakes on Guam. Island invasives: scaling up to meet the challenge. IUCN, Gland, Switzerland, 348–355.
- Siers SR, Shiels AB, Barnhart PD (2020) Invasive snake activity before and after automated aerial baiting. The Journal of Wildlife Management 84(2): 256–267. https://doi.org/10.1002/jwmg.21794
- Sih A, Bell A, Johnson JC (2004) Behavioral syndromes: An ecological and evolutionary overview. Trends in Ecology & Evolution 19(7): 372–378. https://doi.org/10.1016/j.tree.2004.04.009
- Sih A, Cote J, Evans M, Fogarty S, Pruitt J (2012) Ecological implications of behavioural syndromes. Ecology Letters 15(3): 278–289. https://doi.org/10.1111/j.1461-0248.2011.01731.x
- Simard M, Zhang K, Rivera-Monroy VH, Ross MS, Ruiz PL, Castañeda-Moya E, Twilley RR, Rodriguez E (2006) Mapping height and biomass of mangrove forests in Everglades National Park with SRTM elevation data. Photogrammetric Engineering and Remote Sensing 72(3): 299–311. https://doi.org/10.14358/PERS.72.3.299
- Skelton J, Bartoszek I, Beaver CE, Hart KM, Hunter ME (2021) Genome-Wide SNP analysis reveals multiple paternity in Burmese pythons invasive to the greater Florida everglades. Journal of Herpetology 55(4): 355–360. https://doi.org/10.1670/20-104
- Smart AS, Tingley R, Weeks AR, Van Rooyen AR, McCarthy MA (2015) Environmental DNA sampling is more sensitive than a traditional survey technique for detecting an aquatic invader. Ecological Applications 25(7): 1944–1952. https://doi.org/10.1890/14-1751.1
- Smith HT, Sementelli A, Meshaka Jr WE, Engeman RM (2007) Reptilian pathogens of the Florida everglades: The associated costs of Burmese pythons. Methods (San Diego, Calif. ) 24: 63–71.
- Smith BJ (2016) Spatial ecology of invasive Burmese pythons in the Florida Everglades (MS thesis, University of Florida).
- Smith BJ, Hart KM, Demopoulos AWJ, Romagosa CM (2019) Greater Everglades Burmese python stable isotope data, 2003–2012, and standard ellipse area literature review, 2018: U.S. Geological Survey data release. https://doi.org/10.5066/P9QQ1KVH [release date: 12/14/2022]
- Smith BJ, Rochford MR, Brien M, Cherkiss MS, Mazzotti FJ, Snow S, Hart KM (2015) Largest breeding aggregation of Burmese pythons (Python bivittatus) Kuhl, 1820 (Squamata: Pythonidae) and Implications for Potential Development of a Control Tool. Reptiles & Amphibians 22(1): 16–19. https://doi.org/10.17161/randa.v22i1.14025
- Smith BJ, Cherkiss MS, Hart KM, Rochford MR, Selby TH, Snow RW, Mazzotti FJ (2016) Betrayal: Radio-tagged Burmese pythons reveal locations of conspecifics in Everglades National Park. Biological Invasions 18(11): 3239–3250. https://doi.org/10.1007/s10530-016-1211-5
- Smith BJ, Hart KM, Mazzotti FJ, Basille M, Romagosa CM (2018) Evaluating GPS biologging technology for studying spatial ecology of large constricting snakes. Animal Biotelemetry 6(1): 1. https://doi.org/10.1186/s40317-018-0145-3
- Smith SN, Jones MD, Marshall BM, Waengsothorn S, Gale GA, Strine CT (2021) Native Burmese pythons exhibit site fidelity and preference for aquatic habitats in an agricultural mosaic. Scientific Reports 11(1): 1–13. https://doi.org/10.1038/s41598-021-86640-1
- Snow RW, Oberhofer L, Mazzotti FJ (2006) Alligator missippiensis (American Alligator). FEEDING. Herpetological Review 37: 80–81.
- Snow RW, Brien ML, Cherkiss MS, Wilkins L, Mazzotti FJ (2007c) Dietary habits of the Burmese python, Python molurus bivittatus, in Everglades National Park, Florida. Herpetological Bulletin, 5–7.
- Snow RW, Johnson VM, Brien ML, Cherkiss MS, Mazzotti FJ (2007b) Python molurus bivittatus (Burmese Python) Nesting. Herpetological Review 38: 93.
- Snow RW, Krysko KL, Enge KM, Oberhofer L, Warren-Bradley A, Wilkins L (2007a) Introduced populations of Boa constrictor (Boidae) and Python molurus bivittatus (Pythonidae) in southern Florida. Biology of Boas and Pythons. Eagle Mouintain Publishing, 416–438.
- Snow RW, Wolf AJ, Greeves BW, Cherkiss MS, Hill R, Mazzotti FJ (2010) Thermoregulation by a Brooding Burmese Python (Python molurus bivittatus) in Florida. Southeastern Naturalist (Steuben, ME) 9(2): 403–405. https://doi.org/10.1656/058.009.0215
- Sorice MG, Donlan CJ (2015) A human-centered framework for innovation in conservation incentive programs. Ambio 44(8): 788–792. https://doi.org/10.1007/s13280-015-0650-z
- Soto-Shoender JR, Gwinn DC, Sovie A, McCleery RA (2020) Life-history traits moderate the susceptibility of native mammals to an invasive predator. Biological Invasions 22(9): 2671–2684. https://doi.org/10.1007/s10530-020-02278-6
- Southwood A, Avens L (2010) Physiological, behavioral, and ecological aspects of migration in reptiles. Journal of Comparative Physiology. B, Biochemical, Systemic, and Environmental Physiology 180(1): 1–23. https://doi.org/10.1007/s00360-009-0415-8
- Sovie AR, McCleery RA, Fletcher Jr RJ, Hart KM (2016) Invasive pythons, not anthropogenic stressors, explain the distribution of a keystone species. Biological Invasions 18(11): 3309–3318. https://doi.org/10.1007/s10530-016-1221-3
- Sparkman AM, Bronikowski AM, Billings JG, Von Borstel D, Arnold SJ (2013) Avian predation and the evolution of life histories in the garter snake Thamnophis elegans. American Midland Naturalist 170(1): 66–85. https://doi.org/10.1674/0003-0031-170.1.66
- Spencer MM, Miller MA, Courtemanche A, Kinsella JM, Funck S (2022) Python bivittatus (Burmese Python). Parasites. Herpetological Review 53: 347–348.
- Stahl RS, Engeman RM, Avery ML, Mauldin RE (2016) Weather constraints on Burmese python survival in the Florida Everglades, USA based on mechanistic bioenergetics estimates of core body temperature. Cogent Biology 2(1): 1239599. https://doi.org/10.1080/23312025.2016.1239599
- Stahlschmidt ZR, DeNardo DF (2010) Parental care in snakes. Reproductive Biology and Phylogeny of Snakes. Science Publishers Inc., Enfield, NH, 673–702. https://doi.org/10.1201/b10879-19
- Stearns SC (1989) Trade-Offs in Life-History Evolution. Functional Ecology 3(3): 259–268. https://doi.org/10.2307/2389364
- Steen DA (2010) Snakes in the grass: Secretive natural histories defy both conventional and progressive statistics. Herpetological Conservation and Biology 5: 183–188. https://doi.org/10.4013/nbc.2010.53.08
- Steen DA, Guyer C, Smith LL (2012) A case study of relative abundance in snakes. Reptile Biodiversity: Standard Methods for Inventory and Monitoring. University of California Press, Berkeley and Los Angeles, California, 287–294.
- Stenglein MD, Jacobson ER, Wozniak EJ, Wellehan JFX, Kincaid A, Gordon M, Porter BF, Baumgartner W, Stahl S, Kelley K, Towner JS, DeRisi JL (2014) Ball python nidovirus: A candidate etiologic agent for severe respiratory disease in Python regius. mBio 5(5): e01484–e14. https://doi.org/10.1128/mBio.01484-14
- Stevenson RD (1985) The relative importance of behavioral and physiological adjustments controlling body temperature in terrestrial ectotherms. American Naturalist 126(3): 362–386. https://doi.org/10.1086/284423
- Stobo-Wilson AM, Murphy BP, Legge SM, Chapple DG, Crawford HM, Dawson SJ, Dickman CR, Doherty TS, Fleming PA, Gentle M, Newsome TM, Palmer R, Rees MW, Ritchie EG, Speed J, Stuart JM, Thompson E, Turpin J, Woinarski JCZ (2021) Reptiles as food: Predation of Australian reptiles by introduced red foxes compounds and complements predation by cats. Wildlife Research 48(5): 470–480. https://doi.org/10.1071/WR20194
- Storey KB, Storey JM (1992) Natural freeze tolerance in ectothermic vertebrates. Annual Review of Physiology 54(1): 619–637. https://doi.org/10.1146/annurev.ph.54.030192.003155
- Sunquist-Blunden C, Montero N (2021) Florida’s endangered species, threatened species, and species of special concern. Florida Fish and Wildlife Conservation Commission. Tallahassee, USA, 1–15. https://myfwc.com/media/1945/threatened-endangered-species.pdf
- Taillie PJ, Hart KM, Sovie AR, McCleery RA (2021) Native mammals lack resilience to invasive generalist predator. Biological Conservation 261: 109290. https://doi.org/10.1016/j.biocon.2021.109290
- Takahara T, Minamoto T, Doi H (2013) Using environmental DNA to estimate the distribution of an invasive fish species in ponds. PLoS ONE 8(2): e56584. https://doi.org/10.1371/journal.pone.0056584
- Taylor D, Katahira L (1988) Radio telemetry as an aid in eradicating remnant feral goats. Wildlife Society Bulletin 16: 297–299.
- Teem JL, Alphey L, Descamps S, Edgington MP, Edwards O, Gemmell N, Harvey-Samuel T, Melnick RL, Oh KP, Piaggio AJ, Saah JR, Schill D, Thomas P, Smith T, Roberts A (2020) Genetic Biocontrol for Invasive Species. Frontiers in Bioengineering and Biotechnology 8: 1–18. https://doi.org/10.3389/fbioe.2020.00452
- Tewes ME, Mock JM, Young JH (2002) Bobcat predation on quail, birds, and mesomammals. National Quail Symposium Proceedings 5: 65–70.
- Tillis SB, Josimovich JM, Miller MA, Hoon-Hanks LL, Hartmann AM, Claunch NM, Iredale ME, Logan TD, Yackel Adams AA, Bartoszek IA, Humphrey JS, Kluever BM, Stenglein MD, Reed RN, Romagosa CM, Wellehan Jr JFX, Ossiboff RJ (2022) Divergent serpentoviruses in free-ranging invasive pythons and native colubrids in southern Florida, United States. Viruses 14(12): 2726. https://doi.org/10.3390/v14122726
- Thompson MB, Speake BK (2003) Energy and nutrient utilisation by embryonic reptiles. Comparative Biochemistry and Physiology. Part A, Molecular & Integrative Physiology 133(3): 529–538. https://doi.org/10.1016/S1095-6433(02)00188-5
- Troisi CA, Mills DS, Wilkinson A, Zulch HE (2019) Behavioral and cognitive factors that affect the success of scent detection dogs. Comparative Cognition & Behavior Reviews 14: 51–76. https://doi.org/10.3819/CCBR.2019.140007
- Tyre AJ, Tenhumberg B, Field SA, Niejalke D, Parris K, Possingham HP (2003) Improving precision and reducing bias in biological surveys: Estimating false-negative error rates. Ecological Applications 13(6): 1790–1801. https://doi.org/10.1890/02-5078
- Tyrrell CL, Christy MT, Rodda GH, Yackel Adams AA, Ellingson AR, Savidge JA, Dean-Bradley K, Bischof R (2009) Evaluation of trap capture in a geographically closed population of brown treesnakes on Guam. Journal of Applied Ecology 46(1): 128–135. https://doi.org/10.1111/j.1365-2664.2008.01591.x
- Udell B, Martin J, Romagosa C, Waddle H, Johnson F, Falk B, Yackel Adams AA, Sarah A, Ketterlin J, Suarez E, Mazzotti F (2022) Open removal models with temporary emigration and population dynamics to inform invasive animal management. Ecology and Evolution 12(8): e9173. https://doi.org/10.1002/ece3.9173
- USFWS (2012) U.S. Fish and Wildlife Service. The Economic Cost of Large Constrictor Snakes. https://www.sprep.org/attachments/VirLib/Global/economic-cost-large-snakes.pdf
- Van Mierop LHS, Barnard SM (1976a) Thermoregulation in a brooding female Python molurus bivittatus (Serpentes: Boidae). Copeia 1976(2): 398–401. https://doi.org/10.2307/1443978
- Van Mierop LHS, Barnard SM (1976b) Observations on the Reproduction of Python molurus bivittatus (Reptilia, Serpentes, Boidae). Journal of Herpetology 10(4): 333–340. https://doi.org/10.2307/1563071
- Vinegar A (1973) The effects of temperature on the growth and development of embryos of the indian python, Python molurus (Reptilia: Serpentes: Boidae). Copeia 1973(1): 171. https://doi.org/10.2307/1442386
- Vishnu SN, Ramesh C, Thirumurugan V, Sathish C (2021) Size matters: first record of minimum male size at maturity and mating of free-ranging, endangered indian python Python molurus. Asian Journal of Conservation Biology 10: 153–158. https://doi.org/10.53562/ajcb.AQOE1932
- Voss RS, Jansa SA (2012) Snake-venom resistance as a mammalian trophic adaptation: Lessons from didelphid marsupials. Biological Reviews of the Cambridge Philosophical Society 87(4): 822–837. https://doi.org/10.1111/j.1469-185X.2012.00222.x
- Wacker S, Skaug HJ, Forseth T, Solem Ø, Ulvan EM, Fiske P, Karlsson S (2021) Considering sampling bias in close‐kin mark-recapture abundance estimates of Atlantic salmon. Ecology and Evolution 11(9): 3917–3932. https://doi.org/10.1002/ece3.7279
- Wagner E (1976) Breeding the Burmese python at Seattle zoo: Python molurus bivittatus. International Zoo Yearbook 16(1): 83–85. https://doi.org/10.1111/j.1748-1090.1976.tb00136.x
- Walden HDS, Iredale ME, Childress A, Wellehan Jr JFX, Ossiboff RJ (2020) Case report: Invasive pentastomes, Raillietiella orientalis (Sambon, 1922), in a Free-Ranging Banded Water Snake (Nerodia fasciata) in North Central Florida, USA. Frontiers in Veterinary Science 7: 1–6. https://doi.org/10.3389/fvets.2020.00467
- Waldron JL, Lanham JD, Bennett SH (2006) Using behaviorally-based seasons to investigate canebrake rattlesnake (Crotalus horridus) movement patterns and habitat selection. Herpetologica 62(4): 389–398. https://doi.org/10.1655/0018-0831(2006)62[389:UBSTIC]2.0.CO;2
- Wall F (1921) Ophidia Taprobanica or the Snakes of Ceylon. H. R. Cottle, Columbo, Ceylon, 636 pp. https://doi.org/10.5962/bhl.title.53694
- Walters TM, Mazzotti FJ, Fitz HC (2016) Habitat selection by the invasive species Burmese python in southern Florida. Journal of Herpetology 50(1): 50–56. https://doi.org/10.1670/14-098
- Ward RJ, Griffiths RA, Wilkinson JW, Cornish N (2017) Optimising monitoring efforts for secretive snakes: A comparison of occupancy and N-mixture models for assessment of population status. Scientific Reports 7(1): 1–12. https://doi.org/10.1038/s41598-017-18343-5
- Weiss JL, Overpeck JT, Strauss B (2011) Implications of recent sea level rise science for low-elevation areas in coastal cities of the conterminous U.S.A: A letter. Climatic Change 105(3-4): 635–645. https://doi.org/10.1007/s10584-011-0024-x
- Westfall AK, Miller MA, Murray CM, Falk BG, Guyer C, Romagosa CM (2019) Host-specific phenotypic variation of a parasite co-introduced with invasive Burmese pythons. PLoS ONE 14(1): 1–10. https://doi.org/10.1371/journal.pone.0209252
- Whitaker PB, Shine R (2003) A radiotelemetric study of movements and shelter-site selection by free-ranging brownsnakes (Pseudonaja textilis, Elapidae). Herpetological Monograph 17(1): 130–144. https://doi.org/10.1655/0733-1347(2003)017[0130:ARSOMA]2.0.CO;2
- White GC (2005) Correcting wildlife counts using detection probabilities. Wildlife Research 32(3): 211–216. https://doi.org/10.1071/WR03123
- White GC, Burnham KP (1999) Program mark: Survival estimation from populations of marked animals. Bird Study 46(sup1): S120–S139. https://doi.org/10.1080/00063659909477239
- Whitney NM, White CF, Smith BJ, Cherkiss MS, Mazzotti FJ, Hart KM (2021) Accelerometry to study fine-scale activity of invasive Burmese pythons (Python bivittatus) in the wild. Animal Biotelemetry 9(1): 2. https://doi.org/10.1186/s40317-020-00227-7
- Wiewel AS, Yackel Adams AA, Rodda GH (2009) Distribution, density, and biomass of introduced small mammals in the southern mariana islands. Pacific Science 63(2): 205–222. https://doi.org/10.2984/049.063.0204
- Williams R, Thomas L (2009) Cost-effective abundance estimation of rare animals: Testing performance of small-boat surveys for killer whales in British Columbia. Biological Conservation 142(7): 1542–1547. https://doi.org/10.1016/j.biocon.2008.12.028
- Williams BK, Conroy MJ, Nichols JD (2002) Analysis and management of animal populations. Academic Press, 817 pp.
- Williams HJ, Taylor LA, Benhamou S, Bijleveld AI, Clay TA, de Grissac S, Demšar U, English HM, Franconi N, Gómez-Laich A, Griffiths RC, Kay WP, Morales JM, Potts JR, Rogerson KF, Rutz C, Spelt A, Trevail AM, Wilson RP, Börger L (2020) Optimizing the use of biologgers for movement ecology research. Journal of Animal Ecology 89(1): 186–206. https://doi.org/10.1111/1365-2656.13094
- Willson JD (2016) Sampling Reptiles. Reptile Ecology and Conservation, 1–49.
- Willson JD (2017) Indirect effects of invasive Burmese pythons on ecosystems in southern Florida. Journal of Applied Ecology 54(4): 1251–1258. https://doi.org/10.1111/1365-2664.12844
- Willson JD, Gibbons JW (2010) Drift fences, coverboards, and other traps. Amphibian Ecology and Conservation: A Handbook of Techniques. Oxford University Press, 1–17.
- Willson JD, Pittman S (2017) Innovative methods for estimating densities and detection probabilities of secretive reptiles including invasive constrictors and rare upland snakes. Final Report prepared for Department of Defense Legacy Resource. https://apps.dtic.mil/sti/pdfs/AD1046415
- Willson JD, Dorcas ME, Snow RW (2011) Identifying plausible scenarios for the establishment of invasive Burmese pythons (Python molurus) in Southern Florida. Biological Invasions 13(7): 1493–1504. https://doi.org/10.1007/s10530-010-9908-3
- Willson JD, Snow RW, Reed RN, Dorcas ME (2014) Python molurus bivittatus (Burmese python). Minimum size at maturity. Herpetological Review 45: 343–344.
- Willson JD, Pittman SE, Beane JC, Tuberville TD (2018) A novel approach for estimating densities of secretive species from road-survey and spatial-movement data. Wildlife Research 45(5): 446–456. https://doi.org/10.1071/WR16175
- Willson JD, Guzy JC, Durso AM (in press) Overcoming low detectability in snake population research: Case Studies from the southeast USA. Strategies for Conservation Success in Herpetology. SSAR Press.
- Wilson LD, Porras L (1983) The ecological impact of man on the South Florida herpetofauna. 1st Edn. Museum of Natural History, 100 pp. https://doi.org/10.5962/bhl.title.8435
- Wolf AJ, Walters TM, Rochford MR, Snow RW, Mazzotti FJ (2016) Incubation temperature and sex ratio of a Python bivittatus (Burmese python) clutch hatched in Everglades National Park, Florida. Southeastern Naturalist (Steuben, ME) 15(sp8): 35–39. https://doi.org/10.1656/058.015.sp803
- Xu T-L, Sun Y-W, Feng X-Y, Zhou X-N, Zheng B (2021) Development of miRNA-based approaches to explore the interruption of mosquito-borne disease transmission. Frontiers in Cellular and Infection Microbiology 11: 665444. https://doi.org/10.3389/fcimb.2021.665444
- Yackel Adams AA, Lardner B, Knox AJ, Reed RN (2018) Inferring the absence of an incipient population during a rapid response for an invasive species. PLoS ONE 13(9): e0204302. https://doi.org/10.1371/journal.pone.0204302
- Yackel Adams AA, Nafus MG, Klug PE, Lardner B, Mazurek MJ, Savidge JA, Reed RN (2019) Contact rates with nesting birds before and after invasive snake removal: Estimating the effects of trap-based control. NeoBiota 49: 1–17. https://doi.org/10.3897/neobiota.49.35592
- Závorka L, Lang I, Raffard A, Evangelista C, Britton JR, Olden JD, Cucherousset J (2018) Importance of harvest-driven trait changes for invasive species management. Frontiers in Ecology and the Environment 16(6): 317–318. https://doi.org/10.1002/fee.1922
- Zipkin EF, Sullivan PJ, Cooch EG, Kraft CE, Shuter BJ, Weidel BC (2008) Overcompensatory response of a smallmouth bass (Micropterus dolomieu) population to harvest: Release from competition? Canadian Journal of Fisheries and Aquatic Sciences 65(10): 2279–2292. https://doi.org/10.1139/F08-133
- Zipkin EF, Kraft CE, Cooch EG, Sullivan PJ (2009) When can efforts to control nuisance and invasive species backfire? Ecological Applications 19(6): 1585–1595. https://doi.org/10.1890/08-1467.1
- Zippen C (1956) An evaluation of the removal method of estimating animal populations. Biometrics 12(2): 163–189. https://doi.org/10.2307/3001759
Supplementary materials
FWC SFWMD mass length
Data type: table (excel file)
Explanation note: Burmese python (Python molurus bivittatus) length (SVL, cm), mass (kg), and sex recorded from 2017–2022 (n = 4,825) by contractors capturing individuals as part of Removal Programs operated by the Florida Fish and Wildlife Conservation Commission (FWC) and South Florida Water Management District (SFWMD; see Removals section). Data owned and managed by FWC and SFWMD.
Python removals WIMS data used in synthesis
Data type: table (excel file)
Explanation note: Reported removals of Burmese pythons (Python molurus bivittatus) across southern Florida through December 31st, 2021 (n=13,746) used to construct Fig.